Discovery and structure-activity relationship of imidazolinylindole derivatives as kallikrein 7 inhibitors.
Murafuji, H., Muto, T., Goto, M., Imajo, S., Sugawara, H., Oyama, Y., Minamitsuji, Y., Miyazaki, S., Murai, K., Fujioka, H.(2019) Bioorg Med Chem Lett 29: 334-338
- PubMed: 30522951 Search on PubMed
- DOI: https://doi.org/10.1016/j.bmcl.2018.11.011
- Primary Citation Related Structures:
6AHS - PubMed Abstract:
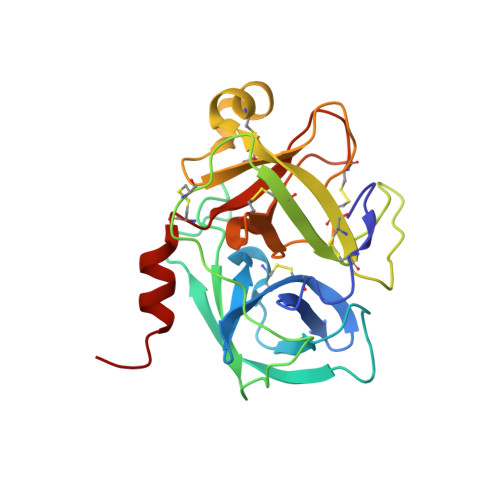
A series of imidazolinylindole derivatives were discovered as novel kallikrein 7 (KLK7, stratum corneum chymotryptic enzyme) inhibitors. Structure-activity relationship (SAR) studies led to the identification of potent human KLK7 inhibitors. By further modification of the benzenesulfonyl moiety to overcome species differences in inhibitory activity, potent inhibitors against both human and mouse KLK7 were identified. Furthermore, the complex structure of 25 with mouse KLK7 could explain the SAR and the cause of the species differences in inhibitory activity.
- Asubio Pharma Co., Ltd, 6-4-3 Minatojima-Minamimachi, Chuo-ku, Kobe 650-0047, Japan.
Organizational Affiliation: