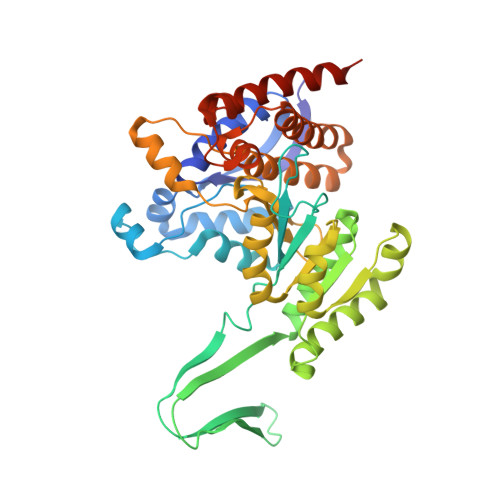
Crystal structures of pan-IDH inhibitor AG-881 in complex with mutant human IDH1 and IDH2
Ma, R., Yun, C.H.(2018) Biochem Biophys Res Commun 503: 2912-2917
- PubMed: 30131249 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2018.08.068
- Primary Citation Related Structures:
6ADG, 6ADI - PubMed Abstract:
Some mutations of isocitrate dehydrogenase 1 and 2 observed in multiple kinds of malignant tumors can lead to a neomorphic enzyme activity that converts alpha-ketoglutarate (α-KG) to 2-hydroxyglutarate (2-HG). As an oncometabolite, 2-HG can cause epigenetic changes and impair cell differentiation. Inhibiting the activity of isocitrate dehydrogenase mutants (mIDH) is considered to be an effective therapy for the treatment of mIDH positive cancers, including glioma and acute myeloid leukemia (AML). The presently disclosed allosteric inhibitors work only on one of the mIDH1 and mIDH2, and it is shown that mIDH1 and mIDH2 have different allosteric inhibition pockets. However, AG-881 from Agios Pharmaceuticals was found to be a pan-IDH inhibitor against both mIDH1 and mIDH2, and is undergoing Phase I clinical trials for tumors with an IDH1 and/or IDH2 mutation. To understand the binding mode of AG-881 to mIDHs, we solved the crystal structures of IDH1-R132H/NADPH/AG-881 and IDH2-R140Q/NADPH/AG-881 complexes, and acquired the IC 50 values of AG-881 for IDH1-R132H and IDH2-R140Q homodimers after different pre-incubation times. Our data show that AG-881 binds IDH1-R132H and IDH2-R140Q in the same allosteric pockets and that the subtle difference in the pockets of these two proteins may contribute to their remarkably different inhibitory kinetics by AG-881. The structural pharmacological data provided in this report may benefit the future development of pan-IDH inhibitors targeting mIDH1 and mIDH2.
- Department of Biophysics, School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, China; Institute of Systems Biomedicine, School of Basic Medical Sciences, Peking University Health Science Center, Beijing 100191, China; Beijing Key Laboratory of Tumor Systems Biology, Peking University Health Science Center, Beijing 100191, China.
Organizational Affiliation: