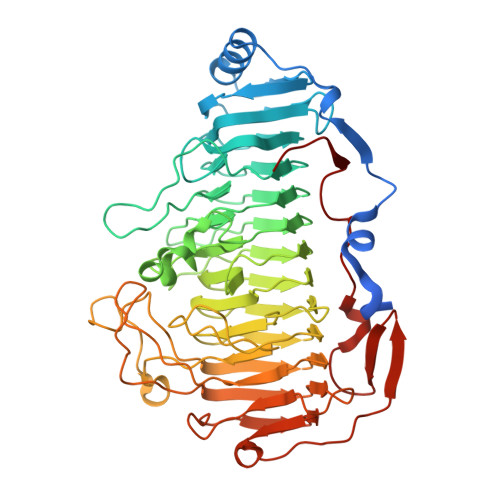
Structural insights into a novel Ca2+-independent PL-6 alginate lyase from Vibrio OU02 identify the possible subsites responsible for product distribution.
Lyu, Q., Zhang, K., Shi, Y., Li, W., Diao, X., Liu, W.(2019) Biochim Biophys Acta Gen Subj 1863: 1167-1176
- PubMed: 31004719 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbagen.2019.04.013
- Primary Citation Related Structures:
5Z9T, 6A40, 6ITG - PubMed Abstract:
Alginate lyases have a wide range of industrial applications, such as oligosaccharide preparation, medical treatment, and bioconversion. Therefore, the discovery and characterization of novel alginate lyases are extremely important. PL-6 alginate lyases are classified into two groups: those with a single domain or two domains. However, only one structure of a two-domain alginate lyase has been determined to date. In this study, we characterized a novel single-domain PL-6 alginate lyase (named AlyF). According to the biochemical analysis, AlyF possesses unique features compared with other PL-6 enzymes, including (1) a Ca 2+ -independent catalytic mechanism and (2) a PolyG-specific cleavage specificity that predominantly produces trisaccharides. The structures of AlyF and its complexes described here reveal the structural basis for these unique features and substrate binding mechanisms, which were further confirmed using mutagenesis. More importantly, we determined the possible subsites specifying the predominantly trisaccharide products of AlyF, which may facilitate the rational design of AlyF for potential applications in preparing a single alginate oligomer.
- MOE Key Laboratory of Marine Genetics and Breeding, College of Marine Life Sciences, Ocean University of China, Qingdao 266003, China; Laboratory for Marine Biology and Biotechnology, Qingdao National Laboratory for Marine Science and Technology, Qingdao 266235, China.
Organizational Affiliation: