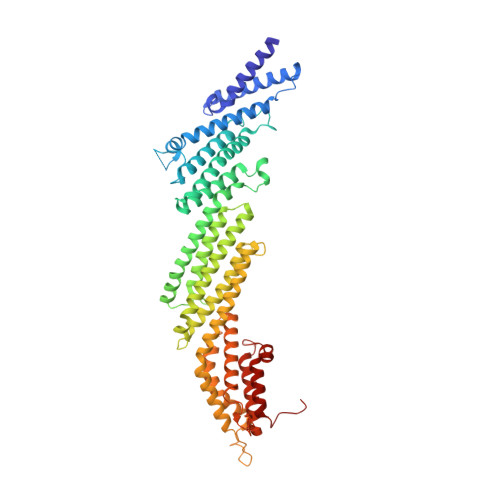
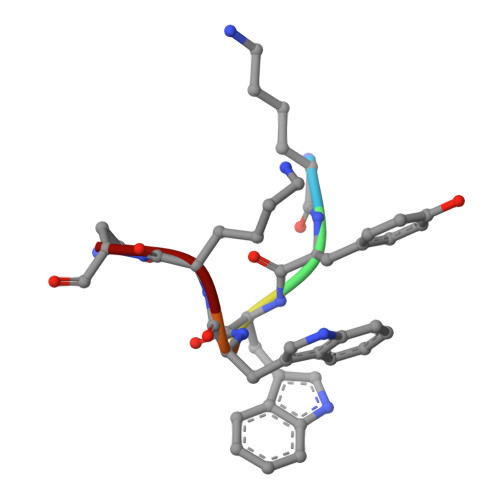
Munc18 and Munc13 serve as a functional template to orchestrate neuronal SNARE complex assembly.
Wang, S., Li, Y., Gong, J.H., Ye, S., Yang, X.F., Zhang, R.G., Ma, C.(2019) Nat Commun 10: 69-69
- PubMed: 30622273 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-018-08028-6
- Primary Citation Related Structures:
6A30 - PubMed Abstract:
The transition of the Munc18-1/syntaxin-1 complex to the SNARE complex, a key step involved in exocytosis, is regulated by Munc13-1, SNAP-25 and synaptobrevin-2, but the underlying mechanism remains elusive. Here, we identify an interaction between Munc13-1 and the membrane-proximal linker region of synaptobrevin-2, and reveal its essential role in transition and exocytosis. Upon this interaction, Munc13-1 not only recruits synaptobrevin-2-embedded vesicles to the target membrane but also renders the synaptobrevin-2 SNARE motif more accessible to the Munc18-1/syntaxin-1 complex. Afterward, the entry of SNAP-25 leads to a half-zippered SNARE assembly, which eventually dissociates the Munc18-1/syntaxin-1 complex to complete SNARE complex formation. Our data suggest that Munc18-1 and Munc13-1 together serve as a functional template to orchestrate SNARE complex assembly.
- Key Laboratory of Molecular Biophysics of the Ministry of Education, College of Life Science and Technology, and the Collaborative Innovation Center for Brain Science, Huazhong University of Science and Technology, 430074, Wuhan, China.
Organizational Affiliation: