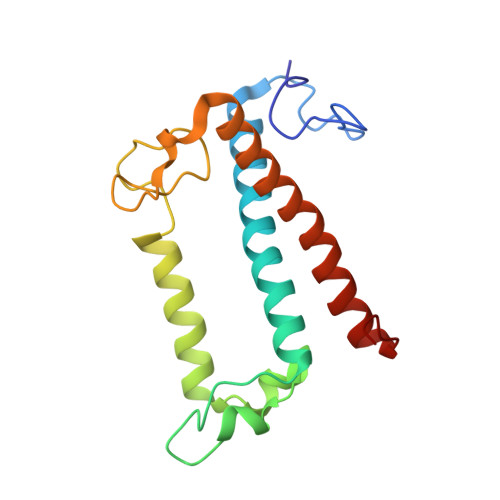
Structural basis for blue-green light harvesting and energy dissipation in diatoms.
Wang, W., Yu, L.J., Xu, C., Tomizaki, T., Zhao, S., Umena, Y., Chen, X., Qin, X., Xin, Y., Suga, M., Han, G., Kuang, T., Shen, J.R.(2019) Science 363
- PubMed: 30733387 Search on PubMed
- DOI: https://doi.org/10.1126/science.aav0365
- Primary Citation Related Structures:
6A2W - PubMed Abstract:
Diatoms are abundant photosynthetic organisms in aquatic environments and contribute 40% of its primary productivity. An important factor that contributes to the success of diatoms is their fucoxanthin chlorophyll a/c-binding proteins (FCPs), which have exceptional light-harvesting and photoprotection capabilities. Here, we report the crystal structure of an FCP from the marine diatom Phaeodactylum tricornutum , which reveals the binding of seven chlorophylls (Chls) a, two Chls c, seven fucoxanthins (Fxs), and probably one diadinoxanthin within the protein scaffold. Efficient energy transfer pathways can be found between Chl a and c, and each Fx is surrounded by Chls, enabling the energy transfer and quenching via Fx highly efficient. The structure provides a basis for elucidating the mechanisms of blue-green light harvesting, energy transfer, and dissipation in diatoms.
- Photosynthesis Research Center, Key Laboratory of Photobiology, Institute of Botany, Chinese Academy of Sciences, 100093 Beijing, China.
Organizational Affiliation: