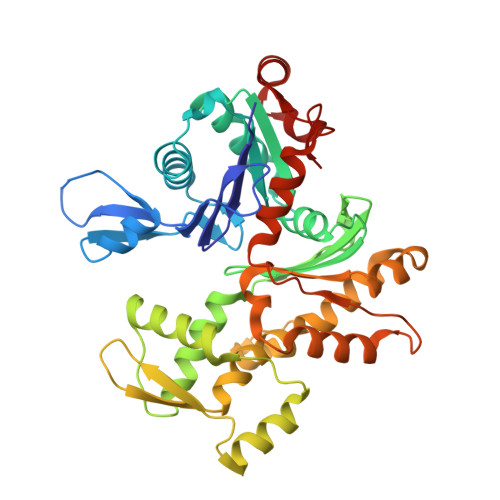
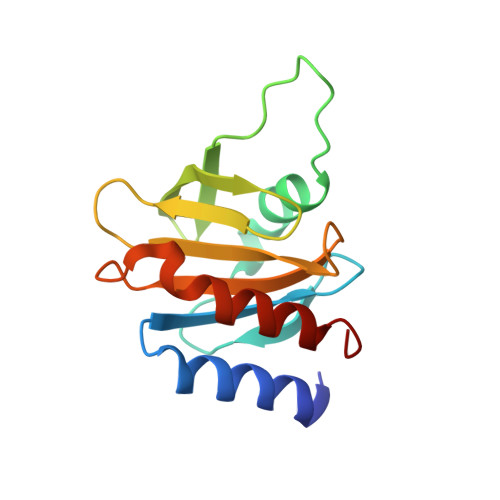
Genomes of Asgard archaea encode profilins that regulate actin.
Akil, C., Robinson, R.C.(2018) Nature 562: 439-443
- PubMed: 30283132 Search on PubMed
- DOI: https://doi.org/10.1038/s41586-018-0548-6
- Primary Citation Related Structures:
5YED, 5YEE, 5ZZA, 5ZZB - PubMed Abstract:
The origin of the eukaryotic cell is unresolved 1,2 . Metagenomics sequencing has recently identified several potential eukaryotic gene homologues in Asgard archaea 3,4 , consistent with the hypothesis that the eukaryotic cell evolved from within the Archaea domain. However, many of these eukaryotic-like sequences are highly divergent and the organisms have yet to be imaged or cultivated, which brings into question the extent to which these archaeal proteins represent functional equivalents of their eukaryotic counterparts. Here we show that Asgard archaea encode functional profilins and thereby establish that this archaeal superphylum has a regulated actin cytoskeleton, one of the hallmarks of the eukaryotic cell 5 . Loki profilin-1, Loki profilin-2 and Odin profilin adopt the typical profilin fold and are able to interact with rabbit actin-an interaction that involves proteins from species that diverged more than 1.2 billion years ago 6 . Biochemical experiments reveal that mammalian actin polymerizes in the presence of Asgard profilins; however, Loki, Odin and Heimdall profilins impede pointed-end elongation. These archaeal profilins also retard the spontaneous nucleation of actin filaments, an effect that is reduced in the presence of phospholipids. Asgard profilins do not interact with polyproline motifs and the profilin-polyproline interaction therefore probably evolved later in the Eukarya lineage. These results suggest that Asgard archaea possess a primordial, polar, profilin-regulated actin system, which may be localized to membranes owing to the sensitivity of Asgard profilins to phospholipids. Because Asgard archaea are also predicted to encode potential eukaryotic-like genes involved in membrane-trafficking and endocytosis 3,4 , imaging is now necessary to elucidate whether these organisms are capable of generating eukaryotic-like membrane dynamics that are regulated by actin, such as are observed in eukaryotic cell movement, podosomes and endocytosis.
- Institute of Molecular and Cell Biology, A*STAR (Agency for Science, Technology and Research), Biopolis, Singapore, Singapore.
Organizational Affiliation: