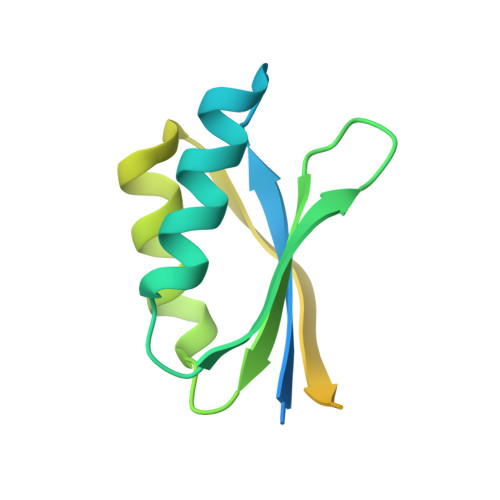
Specific recognition of two MAX effectors by integrated HMA domains in plant immune receptors involves distinct binding surfaces
Guo, L., Cesari, S., de Guillen, K., Chalvon, V., Mammri, L., Ma, M., Meusnier, I., Bonnot, F., Padilla, A., Peng, Y.L., Liu, J., Kroj, T.(2018) Proc Natl Acad Sci U S A 115: 11637-11642
- PubMed: 30355769 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1810705115
- Primary Citation Related Structures:
5ZNE, 5ZNG - PubMed Abstract:
The structurally conserved but sequence-unrelated MAX ( Magnaporthe oryzae avirulence and ToxB-like) effectors AVR1-CO39 and AVR-PikD from the blast fungus M. oryzae are recognized by the rice nucleotide-binding domain and leucine-rich repeat proteins (NLRs) RGA5 and Pikp-1, respectively. This involves, in both cases, direct interaction of the effector with a heavy metal-associated (HMA) integrated domain (ID) in the NLR. Here, we solved the crystal structures of a C-terminal fragment of RGA5 carrying the HMA ID (RGA5_S), alone, and in complex with AVR1-CO39 and compared it to the structure of the Pikp1 HMA /AVR-PikD complex. In both complexes, HMA ID/MAX effector interactions involve antiparallel alignment of β-sheets from each partner. However, effector-binding occurs at different surfaces in Pikp1 HMA and RGA5 HMA , indicating that these interactions evolved independently by convergence of these two MAX effectors to the same type of plant target proteins. Interestingly, the effector-binding surface in RGA5 HMA overlaps with the surface that mediates RGA5 HMA self-interaction. Mutations in the HMA-binding interface of AVR1-CO39 perturb RGA5 HMA -binding, in vitro and in vivo, and affect the recognition of M. oryzae in a rice cultivar containing Pi-CO39 Our study provides detailed insight into the mechanisms of effector recognition by NLRs, which has substantial implications for future engineering of NLRs to expand their recognition specificities. In addition, we propose, as a hypothesis for the understanding of effector diversity, that in the structurally conserved MAX effectors the molecular mechanism of host target protein-binding is conserved rather than the host target proteins themselves.
- State Key Laboratory of Agrobiotechnology, China Agricultural University, 100083 Beijing, People's Republic of China.
Organizational Affiliation: