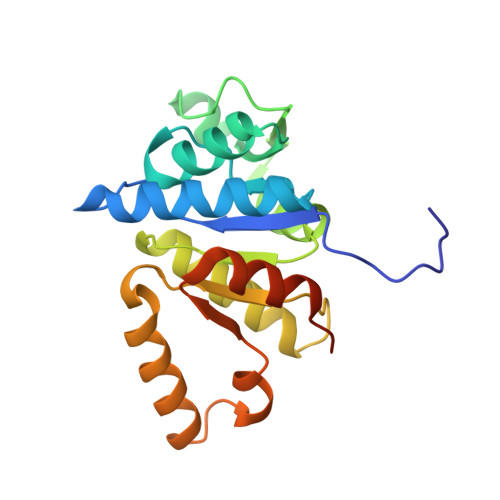
Structural insight into molecular mechanism of cytokinin activating protein from Pseudomonas aeruginosa PAO1.
Seo, H., Kim, K.J.(2018) Environ Microbiol 20: 3214-3223
- PubMed: 29901273 Search on PubMed
- DOI: https://doi.org/10.1111/1462-2920.14287
- Primary Citation Related Structures:
5ZBJ, 5ZBK, 5ZBL - PubMed Abstract:
Cytokinin (CK)-activating enzyme, called LOG, is a phosphoribohydrolase that hydrolyzes nucleotides into nucleobases and phosphoriboses. This reaction is a fascinating target for regulation of cellular active CK. However, misannotation of LOG as a lysine decarboxylase and the lack of detailed catalytic and substrate-binding mechanisms have prevented studies of LOG at a protein-level. In this study, we determined the crystal structure of PA4923 from Pseudomonas aeruginosa PAO1. The overall structure of PA4923 resembles those of type-I LOGs, and it exhibited phosphoribohydrolase activity against AMP. These observations indicated that PA4923 functions as an LOG. We also determined the PaLOG structure in complex with AMP and elucidated the detailed binding mode of LOG against the AMP substrate. Interestingly, PaLOG undergoes an open/closed conformational change upon binding AMP, during which the Glu74 residue located on the β3-β4 connecting loop flips 180° and moves 13 Å towards the AMP molecule. Structural and amino acid sequence comparisons of LOGs suggest that this conformational change upon substrate binding might be a common phenomenon in LOGs. In addition, based on our structural studies and the reported catalytic mechanism of nucleoside hydrolases, we proposed a catalytic mechanism for LOG in which an oxocarbenium ion-like transition state is formed.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daegu, 41566, Republic of Korea.
Organizational Affiliation: