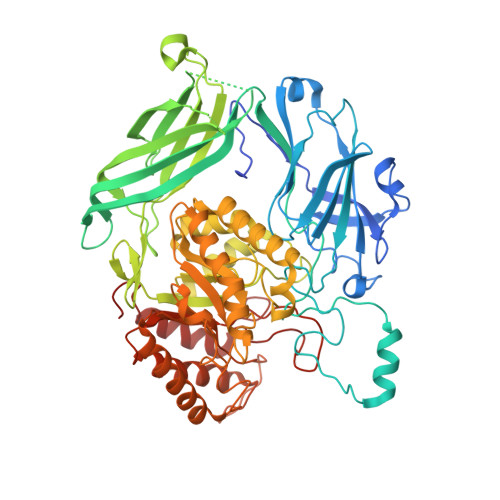
Dissection of the substrate preference and structure of gut microbial-glucuronidases identifies the major bacteria causing xenobiotic toxicity
Dashnyam, P., Mudududdla, R., Hsieh, T.J., Lin, T.C., Lin, H.Y., Chen, P.Y., Hsu, C.Y., Lin, C.H.To be published.