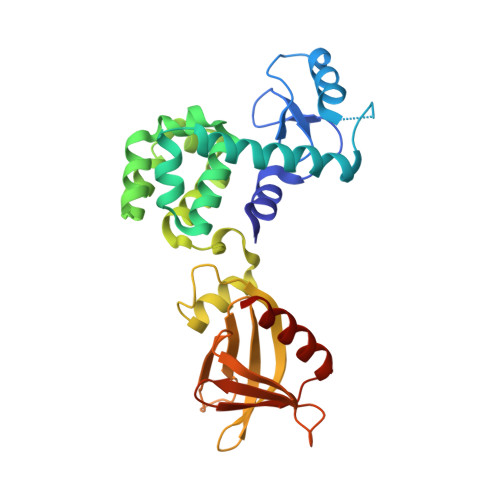
Structural basis of sterol recognition and nonvesicular transport by lipid transfer proteins anchored at membrane contact sites
Tong, J., Manik, M.K., Im, Y.J.(2018) Proc Natl Acad Sci U S A 115: E856-E865
- PubMed: 29339490
- DOI: https://doi.org/10.1073/pnas.1719709115
- Primary Citation Related Structures:
5YQI, 5YQJ, 5YQP, 5YQQ, 5YQR, 5YS0 - PubMed Abstract:
Membrane contact sites (MCSs) in eukaryotic cells are hotspots for lipid exchange, which is essential for many biological functions, including regulation of membrane properties and protein trafficking. Lipid transfer proteins anchored at membrane contact sites (LAMs) contain sterol-specific lipid transfer domains [StARkin domain (SD)] and multiple targeting modules to specific membrane organelles. Elucidating the structural mechanisms of targeting and ligand recognition by LAMs is important for understanding the interorganelle communication and exchange at MCSs. Here, we determined the crystal structures of the yeast Lam6 pleckstrin homology (PH)-like domain and the SDs of Lam2 and Lam4 in the apo form and in complex with ergosterol. The Lam6 PH-like domain displays a unique PH domain fold with a conserved N-terminal α-helix. The Lam6 PH-like domain lacks the basic surface for phosphoinositide binding, but contains hydrophobic patches on its surface, which are critical for targeting to endoplasmic reticulum (ER)-mitochondrial contacts. Structures of the LAM SDs display a helix-grip fold with a hydrophobic cavity and a flexible Ω1-loop as a lid. Ergosterol is bound to the pocket in a head-down orientation, with its hydrophobic acyl group located in the tunnel entrance. The Ω1-loop in an open conformation is essential for ergosterol binding by direct hydrophobic interaction. Structural comparison suggested that the sterol binding mode of the Lam2 SD2 is likely conserved among the sterol transfer proteins of the StARkin superfamily. Structural models of full-length Lam2 correlated with the sterol transport function at the membrane contact sites.
- College of Pharmacy, Chonnam National University, Bukgu, Gwangju, 61186, Republic of Korea.
Organizational Affiliation: