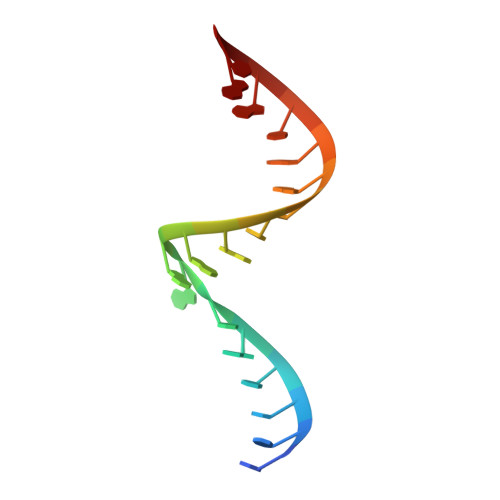
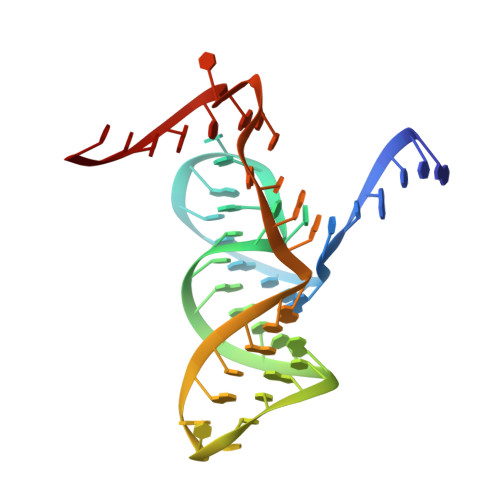
Structure-based insights into self-cleavage by a four-way junctional twister-sister ribozyme
Zheng, L., Mairhofer, E., Teplova, M., Zhang, Y., Ma, J., Patel, D.J., Micura, R., Ren, A.(2017) Nat Commun 8: 1180-1180
- PubMed: 29081514
- DOI: https://doi.org/10.1038/s41467-017-01276-y
- Primary Citation Related Structures:
5Y85, 5Y87 - PubMed Abstract:
Here we report on the crystal structure and cleavage assays of a four-way junctional twister-sister self-cleaving ribozyme. Notably, 11 conserved spatially separated loop nucleotides are brought into close proximity at the ribozyme core through long-range interactions mediated by hydrated Mg 2+ cations. The C62-A63 step at the cleavage site adopts a splayed-apart orientation, with flexible C62 directed outwards, whereas A63 is directed inwards and anchored by stacking and hydrogen-bonding interactions. Structure-guided studies of key base, sugar, and phosphate mutations in the twister-sister ribozyme, suggest contributions to the cleavage chemistry from interactions between a guanine at the active site and the non-bridging oxygen of the scissile phosphate, a feature found previously also for the related twister ribozyme. Our four-way junctional pre-catalytic structure differs significantly in the alignment at the cleavage step (splayed-apart vs. base-stacked) and surrounding residues and hydrated Mg 2+ ions relative to a reported three-way junctional pre-catalytic structure of the twister-sister ribozyme.
- Life Sciences Institute, Zhejiang University, 310058, Hangzhou, China.
Organizational Affiliation: