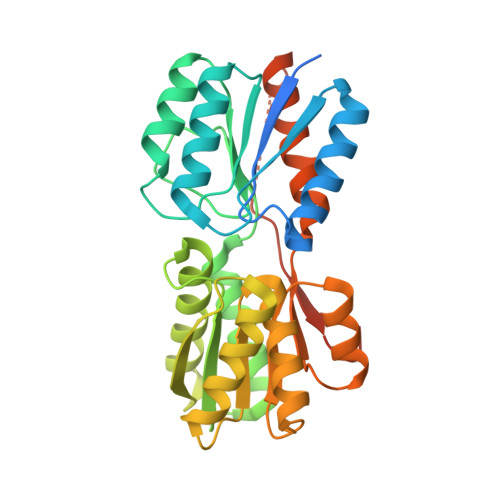
Molecular mechanism of environmental d-xylose perception by a XylFII-LytS complex in bacteria
Li, J., Wang, C., Yang, G., Sun, Z., Guo, H., Shao, K., Gu, Y., Jiang, W., Zhang, P.(2017) Proc Natl Acad Sci U S A 114: 8235-8240
- PubMed: 28716923
- DOI: https://doi.org/10.1073/pnas.1620183114
- Primary Citation Related Structures:
5XSD, 5XSJ, 5XSS - PubMed Abstract:
d-xylose, the main building block of plant biomass, is a pentose sugar that can be used by bacteria as a carbon source for bio-based fuel and chemical production through fermentation. In bacteria, the first step for d-xylose metabolism is signal perception at the membrane. We previously identified a three-component system in Firmicutes bacteria comprising a membrane-associated sensor protein (XylFII), a transmembrane histidine kinase (LytS) for periplasmic d-xylose sensing, and a cytoplasmic response regulator (YesN) that activates the transcription of the target ABC transporter xylFGH genes to promote the uptake of d-xylose. The molecular mechanism underlying signal perception and integration of these processes remains elusive, however. Here we purified the N-terminal periplasmic domain of LytS (LytSN) in a complex with XylFII and determined the conformational structures of the complex in its d-xylose-free and d-xylose-bound forms. LytSN contains a four-helix bundle, and XylFII contains two Rossmann fold-like globular domains with a xylose-binding cleft between them. In the absence of d-xylose, LytSN and XylFII formed a heterodimer. Specific binding of d-xylose to the cleft of XylFII induced a large conformational change that closed the cleft and brought the globular domains closer together. This conformational change led to the formation of an active XylFII-LytSN heterotetramer. Mutations at the d-xylose binding site and the heterotetramer interface diminished heterotetramer formation and impaired the d-xylose-sensing function of XylFII-LytS. Based on these data, we propose a working model of XylFII-LytS that provides a molecular basis for d-xylose utilization and metabolic modification in bacteria.
- National Key Laboratory of Plant Molecular Genetics, Chinese Academy of Sciences Center for Excellence in Molecular Plant Sciences, Institute of Plant Physiology and Ecology, Shanghai Institutes for Biological Sciences, Chinese Academy of Sciences, Shanghai 200032, China.
Organizational Affiliation: