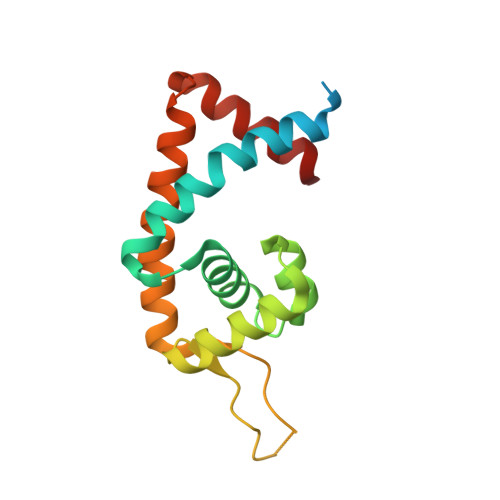
Structural analysis of the regulatory mechanism of MarR protein Rv2887 in M. tuberculosis
Gao, Y.R., Li, D.F., Fleming, J., Zhou, Y.F., Liu, Y., Deng, J.Y., Zhou, L., Zhou, J., Zhu, G.F., Zhang, X.E., Wang, D.C., Bi, L.J.(2017) Sci Rep 7: 6471-6471
- PubMed: 28743871 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41598-017-01705-4
- Primary Citation Related Structures:
5HSM, 5HSO, 5X7Z, 5X80 - PubMed Abstract:
MarR family proteins are transcriptional regulators that control expression of bacterial proteins involved in metabolism, virulence, stress responses and multi-drug resistance, mainly via ligand-mediated attenuation of DNA binding. Greater understanding of their underlying regulatory mechanism may open up new avenues for the effective treatment of bacterial infections. To gain molecular insight into the mechanism of Rv2887, a MarR family protein in M. tuberculosis, we first showed that it binds salicylate (SA) and para-aminosalicylic acid (PAS), its structural analogue and an antitubercular drug, in a 1:1 stoichiometry with high affinity. Subsequent determination and analysis of Rv2887 crystal structures in apo form, and in complex with SA, PAS and DNA showed that SA and PAS bind to Rv2887 at similar sites, and that Rv2887 interacts with DNA mainly by insertion of helix α4 into the major groove. Ligand binding triggers rotation of the wHTH domain of Rv2887 toward the dimerization domain, causing changes in protein conformation such that it can no longer bind to a 27 bp recognition sequence in the upstream region of gene Rv0560c. The structures provided here lay a foundation for the design of small molecules that target Rv2887, a potential new approach for the development of anti-mycobacterials.
- School of Stomatology and Medicine, Foshan University, Foshan, 528000, Guangdong Province, China.
Organizational Affiliation: