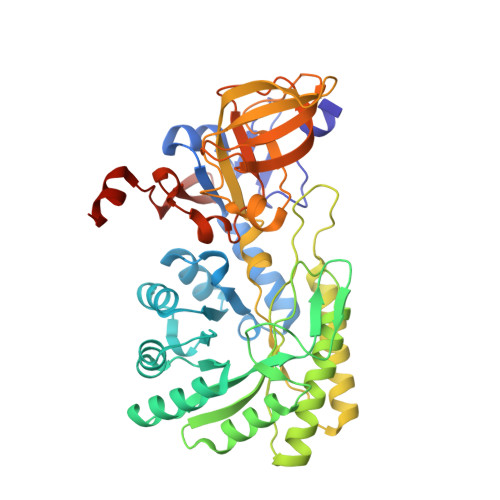
Structural basis for substrate specificity of meso-diaminopimelic acid decarboxylase from Corynebacterium glutamicum.
Son, H.F., Kim, K.J.(2018) Biochem Biophys Res Commun 495: 1815-1821
- PubMed: 29233695
- DOI: https://doi.org/10.1016/j.bbrc.2017.11.097
- Primary Citation of Related Structures:
5X7M, 5X7N - PubMed Abstract:
l-lysine is an essential amino acid that is widely used as a food supplement for humans and animals. meso-Diaminopimelic acid decarboxylase (DAPDC) catalyzes the final step in the de novol-lysine biosynthetic pathway by converting meso-diaminopimelic acid (meso-DAP) into l-lysine by decarboxylation reaction. To elucidate its molecular mechanisms, we determined the crystal structure of DAPDC from Corynebacterium glutamicum (CgDAPDC). The PLP cofactor is bound at the center of the barrel domain and forms a Schiff base with the catalytic Lys75 residue. We also determined the CgDAPDC structure in complex with both pyridoxal 5'-phosphate (PLP) and the l-lysine product and revealed that the protein has an optimal substrate binding pocket to accommodate meso-DAP as a substrate. Structural comparison of CgDAPDC with other amino acid decarboxylases with different substrate specificities revealed that the position of the α15 helix in CgDAPDC and the residues located on the helix are crucial for determining the substrate specificities of the amino acid decarboxylases.
- School of Life Sciences, KNU Creative BioResearch Group, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu, 41566, Republic of Korea; KNU Institute for Microorganisms, Kyungpook National University, Daehak-ro 80, Buk-ku, Daegu, 41566, Republic of Korea.
Organizational Affiliation: