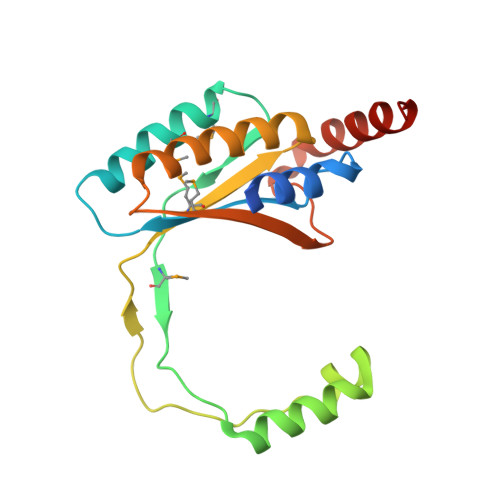
Crystal Structure of the DNA-Binding Domain of Human Herpesvirus 6A Immediate Early Protein 2.
Nishimura, M., Wang, J., Wakata, A., Sakamoto, K., Mori, Y.(2017) J Virol 91
- PubMed: 28794035
- DOI: https://doi.org/10.1128/JVI.01121-17
- Primary Citation of Related Structures:
5WX8 - PubMed Abstract:
Immediate early proteins of human herpesvirus 6A (HHV-6A) are expressed at the outset of lytic infection and thereby regulate viral gene expression. Immediate early protein 2 (IE2) of HHV-6A is a transactivator that drives a variety of promoters. The C-terminal region of HHV-6A IE2 is shared among IE2 homologs in betaherpesviruses and is involved in dimerization, DNA binding, and transcription factor binding. In this study, the structure of the IE2 C-terminal domain (IE2-CTD) was determined by X-ray crystallography at a resolution of 2.5 Å. IE2-CTD forms a homodimer stabilized by a β-barrel core with two interchanging long loops. Unexpectedly, the core structure resembles those of the gammaherpesvirus factors EBNA1 of Epstein-Barr virus and LANA of Kaposi sarcoma-associated herpesvirus, but the interchanging loops are longer in IE2-CTD and form helix-turn-helix (HTH)-like motifs at their tips. The HTH and surrounding α-helices form a structural feature specific to the IE2 group. The apparent DNA-binding site (based on structural similarity with EBNA1 and LANA) resides on the opposite side of the HTH-like motifs, surrounded by positive electrostatic potential. Mapping analysis of conserved residues on the three-dimensional structure delineated a potential factor-binding site adjacent to the expected DNA-binding site. The predicted bi- or tripartite functional sites indicate a role for IE2-CTD as an adapter connecting the promoter and transcriptional factors that drive gene expression. IMPORTANCE Human herpesvirus 6A (HHV-6A) and HHV-6B belong to betaherpesvirus subfamily. Both viruses establish lifelong latency after primary infection, and their reactivation poses a significant risk to immunocompromised patients. Immediate early protein 2 (IE2) of HHV-6A and HHV-6B is a transactivator that triggers viral replication and contains a DNA-binding domain shared with other betaherpesviruses such as human herpesvirus 7 and human cytomegalovirus. In this study, an atomic structure of the DNA-binding domain of HHV-6A IE2 was determined and analyzed, enabling a structure-based understanding of the functions of IE2, specifically DNA recognition and interaction with transcription factors. Unexpectedly, the dimeric core resembles the DNA-binding domain of transcription regulators from gammaherpesviruses, showing structural conservation as a DNA-binding domain but with its own unique structural features. These findings facilitate further characterization of this key viral transactivator.
- Division of Clinical Virology, Kobe University Graduate School of Medicine, Kobe, Japan.
Organizational Affiliation: