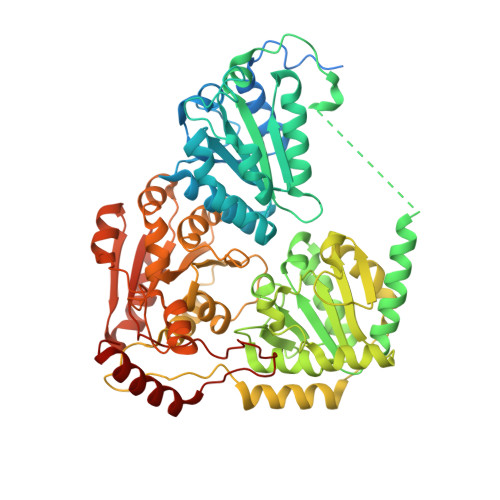
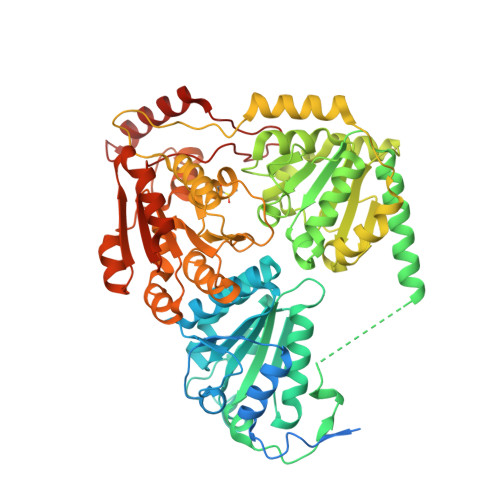
Structural insights into the mechanism of inhibition of AHAS by herbicides.
Lonhienne, T., Garcia, M.D., Pierens, G., Mobli, M., Nouwens, A., Guddat, L.W.(2018) Proc Natl Acad Sci U S A 115: E1945-E1954
- PubMed: 29440497
- DOI: https://doi.org/10.1073/pnas.1714392115
- Primary Citation of Related Structures:
5WJ1, 5WKC - PubMed Abstract:
Acetohydroxyacid synthase (AHAS), the first enzyme in the branched amino acid biosynthesis pathway, is present only in plants and microorganisms, and it is the target of >50 commercial herbicides. Penoxsulam (PS), which is a highly effective broad-spectrum AHAS-inhibiting herbicide, is used extensively to control weed growth in rice crops. However, the molecular basis for its inhibition of AHAS is poorly understood. This is despite the availability of structural data for all other classes of AHAS-inhibiting herbicides. Here, crystallographic data for Saccharomyces cerevisiae AHAS (2.3 Å) and Arabidopsis thaliana AHAS (2.5 Å) in complex with PS reveal the extraordinary molecular mechanisms that underpin its inhibitory activity. The structures show that inhibition of AHAS by PS triggers expulsion of two molecules of oxygen bound in the active site, releasing them as substrates for an oxygenase side reaction of the enzyme. The structures also show that PS either stabilizes the thiamin diphosphate (ThDP)-peracetate adduct, a product of this oxygenase reaction, or traps within the active site an intact molecule of peracetate in the presence of a degraded form of ThDP: thiamine aminoethenethiol diphosphate. Kinetic analysis shows that PS inhibits AHAS by a combination of events involving FAD oxidation and chemical alteration of ThDP. With the emergence of increasing levels of resistance toward front-line herbicides and the need to optimize the use of arable land, these data suggest strategies for next generation herbicide design.
- School of Chemistry and Molecular Biosciences, The University of Queensland, Brisbane, QLD 4072, Australia; t.lonhienne@uq.edu.au luke.guddat@uq.edu.au.
Organizational Affiliation: