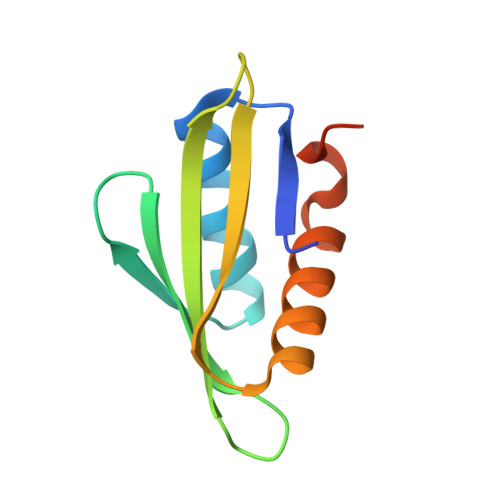
Intramolecular autoinhibition of checkpoint kinase 1 is mediated by conserved basic motifs of the C-terminal kinase-associated 1 domain.
Emptage, R.P., Schoenberger, M.J., Ferguson, K.M., Marmorstein, R.(2017) J Biological Chem 292: 19024-19033
- PubMed: 28972186 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M117.811265
- Primary Citation Related Structures:
5WI2 - PubMed Abstract:
Precise control of the cell cycle allows for timely repair of genetic material prior to replication. One factor intimately involved in this process is checkpoint kinase 1 (Chk1), a DNA damage repair inducing Ser/Thr protein kinase that contains an N-terminal kinase domain and a C-terminal regulatory region consisting of a ∼100-residue linker followed by a putative kinase-associated 1 (KA1) domain. We report the crystal structure of the human Chk1 KA1 domain, demonstrating striking structural homology with other sequentially diverse KA1 domains. Separately purified Chk1 kinase and KA1 domains are intimately associated in solution, which results in inhibition of Chk1 kinase activity. Using truncation mutants and site-directed mutagenesis, we define the inhibitory face of the KA1 domain as a series of basic residues residing on two conserved regions of the primary structure. These findings point to KA1-mediated intramolecular autoinhibition as a key regulatory mechanism of human Chk1, and provide new therapeutic possibilities with which to attack this validated oncology target with small molecules.
- From the Department of Biochemistry and Biophysics and the Abramson Family Cancer Research Institute, Perelman School of Medicine, University of Pennsylvania, Philadelphia, Pennsylvania 19104, emptage@upenn.edu.
Organizational Affiliation: