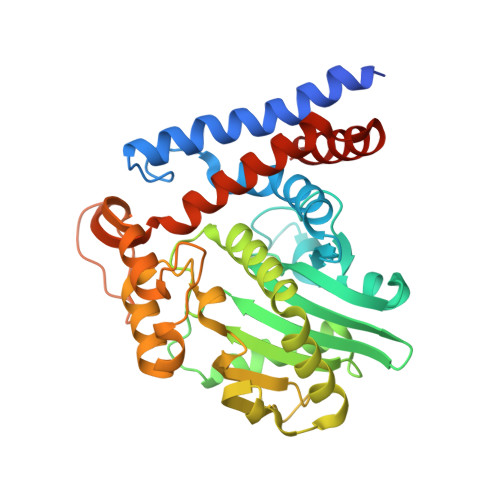
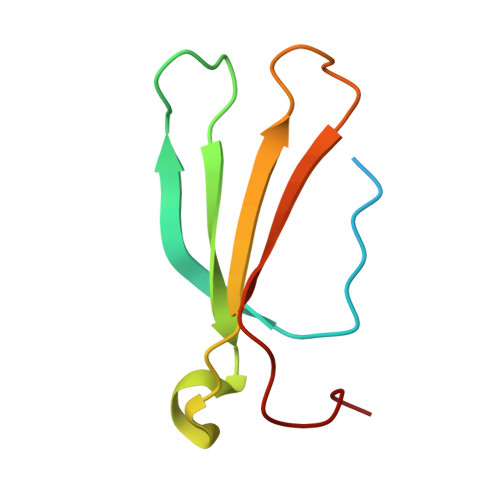
Mechanism of host substrate acetylation by a YopJ family effector.
Zhang, Z.M., Ma, K.W., Gao, L., Hu, Z., Schwizer, S., Ma, W., Song, J.(2017) Nat Plants 3: 17115-17115
- PubMed: 28737762 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nplants.2017.115
- Primary Citation Related Structures:
5W3T, 5W3X, 5W3Y, 5W40 - PubMed Abstract:
The Yersinia outer protein J (YopJ) family of bacterial effectors depends on a novel acetyltransferase domain to acetylate signalling proteins from plant and animal hosts. However, the underlying mechanism is unclear. Here, we report the crystal structures of PopP2, a YopJ effector produced by the plant pathogen Ralstonia solanacearum, in complex with inositol hexaphosphate (InsP 6 ), acetyl-coenzyme A (AcCoA) and/or substrate Resistance to Ralstonia solanacearum 1 (RRS1-R) WRKY . PopP2 recognizes the WRKYGQK motif of RRS1-R WRKY to position a targeted lysine in the active site for acetylation. Importantly, the PopP2-RRS1-R WRKY association is allosterically regulated by InsP 6 binding, suggesting a previously unidentified role of the eukaryote-specific cofactor in substrate interaction. Furthermore, we provide evidence for the reaction intermediate of PopP2-mediated acetylation, an acetyl-cysteine covalent adduct, lending direct support to the 'ping-pong'-like catalytic mechanism proposed for YopJ effectors. Our study provides critical mechanistic insights into the virulence activity of YopJ class of acetyltransferases.
- Department of Biochemistry, University of California, Riverside, California 92521, USA.
Organizational Affiliation: