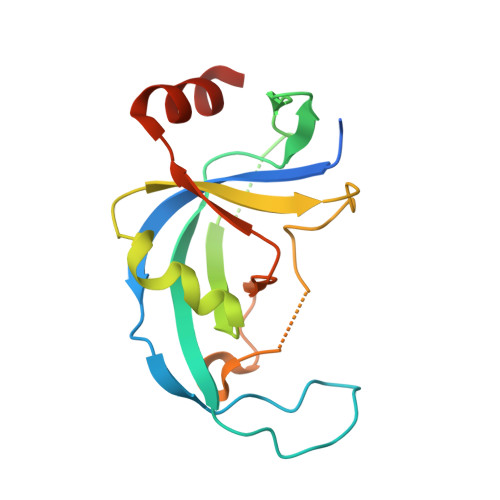
Structural and functional analysis of an OB-fold in human Ctc1 implicated in telomere maintenance and bone marrow syndromes.
Shastrula, P.K., Rice, C.T., Wang, Z., Lieberman, P.M., Skordalakes, E.(2018) Nucleic Acids Res 46: 972-984
- PubMed: 29228254 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkx1213
- Primary Citation Related Structures:
5W2L - PubMed Abstract:
The human CST (Ctc1, Stn1 and Ten1) complex binds the telomeric overhang and regulates telomere length by promoting C-strand replication and inhibiting telomerase-dependent G-strand synthesis. Structural and biochemical studies on the human Stn1 and Ten1 complex revealed its mechanism of assembly and nucleic acid binding. However, little is known about the structural organization of the multi-domain Ctc1 protein and how each of these domains contribute to telomere length regulation. Here, we report the structure of a central domain of human Ctc1. The structure reveals a canonical OB-fold with the two identified disease mutations (R840W and V871M) contributing to the fold of the protein. In vitro assays suggest that although this domain is not contributing directly to Ctc1's substrate binding properties, it affects full-length Ctc1 localization to telomeres and Stn1-Ten1 binding. Moreover, functional assays show that deletion of the entire OB-fold domain leads to significant increase in telomere length, frequency of internal single G-strands and fragile telomeres. Our findings demonstrate that a previously unknown OB-fold domain contributes to efficient Ctc1 telomere localization and chromosome end maintenance.
- The Wistar Institute, Gene expression and regulation program, 3601 Spruce Street, Philadelphia, PA 19104, USA.
Organizational Affiliation: