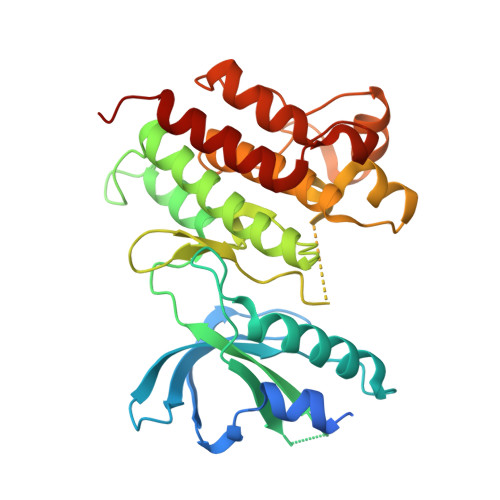
The Axl kinase domain in complex with a macrocyclic inhibitor offers first structural insights into an active TAM receptor kinase.
Gajiwala, K.S., Grodsky, N., Bolanos, B., Feng, J., Ferre, R., Timofeevski, S., Xu, M., Murray, B.W., Johnson, T.W., Stewart, A.(2017) J Biological Chem 292: 15705-15716
- PubMed: 28724631 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.771485
- Primary Citation Related Structures:
5U6B, 5U6C - PubMed Abstract:
The receptor tyrosine kinase family consisting of Tyro3, Axl, and Mer (TAM) is one of the most recently identified receptor tyrosine kinase families. TAM receptors are up-regulated postnatally and maintained at high levels in adults. They all play an important role in immunity, but Axl has also been implicated in cancer and therefore is a target in the discovery and development of novel therapeutics. However, of the three members of the TAM family, the Axl kinase domain is the only one that has so far eluded structure determination. To this end, using differential scanning fluorimetry and hydrogen-deuterium exchange mass spectrometry, we show here that a lower stability and greater dynamic nature of the Axl kinase domain may account for its poor crystallizability. We present the first structural characterization of the Axl kinase domain in complex with a small-molecule macrocyclic inhibitor. The Axl crystal structure revealed two distinct conformational states of the enzyme, providing a first glimpse of what an active TAM receptor kinase may look like and suggesting a potential role for the juxtamembrane region in enzyme activity. We noted that the ATP/inhibitor-binding sites of the TAM members closely resemble each other, posing a challenge for the design of a selective inhibitor. We propose that the differences in the conformational dynamics among the TAM family members could potentially be exploited to achieve inhibitor selectivity for targeted receptors.
- From Oncology Medicinal Chemistry and ketan.gajiwala@pfizer.com.
Organizational Affiliation: