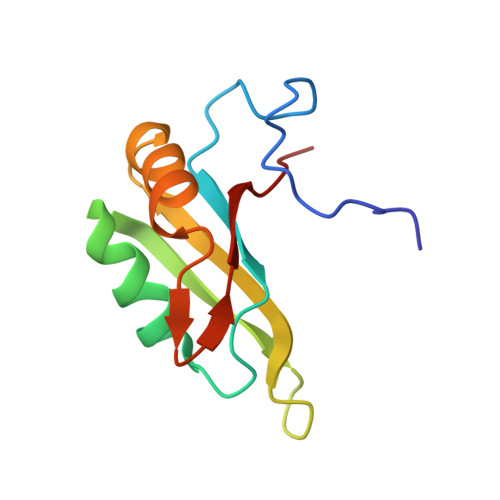
Oligomeric transition and dynamics of RNA binding by the HuR RRM1 domain in solution.
Lixa, C., Mujo, A., de Magalhaes, M.T.Q., Almeida, F.C.L., Lima, L.M.T.R., Pinheiro, A.S.(2018) J Biomol NMR 72: 179-192
- PubMed: 30535889 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-018-0217-y
- Primary Citation Related Structures:
5SZW - PubMed Abstract:
Human antigen R (HuR) functions as a major post-transcriptional regulator of gene expression through its RNA-binding activity. HuR is composed by three RNA recognition motifs, namely RRM1, RRM2, and RRM3. The two N-terminal RRM domains are disposed in tandem and contribute mostly to HuR interaction with adenine and uracil-rich elements (ARE) in mRNA. Here, we used a combination of NMR and electrospray ionization-ion mobility spectrometry-mass spectrometry (ESI-IMS-MS) to characterize the structure, dynamics, RNA recognition, and dimerization of HuR RRM1. Our solution structure reveals a canonical RRM fold containing a 19-residue, intrinsically disordered N-terminal extension, which is not involved in RNA binding. NMR titration results confirm the primary RNA-binding site to the two central β-strands, β1 and β3, for a cyclooxygenase 2 (Cox2) ARE I-derived, 7-nucleotide RNA ligand. We show by 15 N relaxation that, in addition to the N- and C-termini, the β2-β3 loop undergoes fast backbone dynamics (ps-ns) both in the free and RNA-bound state, indicating that no structural ordering happens upon RNA interaction. ESI-IMS-MS reveals that HuR RRM1 dimerizes, however dimer population represents a minority. Dimerization occurs via the α-helical surface, which is oppositely orientated to the RNA-binding β-sheet. By using a DNA analog of the Cox2 ARE I, we show that DNA binding stabilizes HuR RRM1 monomer and shifts the monomer-dimer equilibrium toward the monomeric species. Altogether, our results deepen the current understanding of the mechanism of RNA recognition employed by HuR.
- Department of Biochemistry, Institute of Chemistry, Federal University of Rio de Janeiro, Rio de Janeiro, 21941-909, Brazil.
Organizational Affiliation: