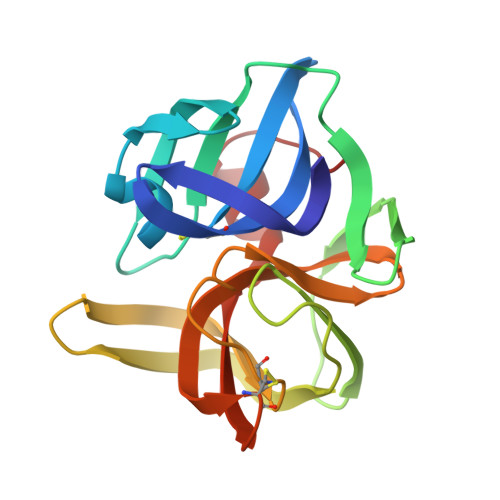
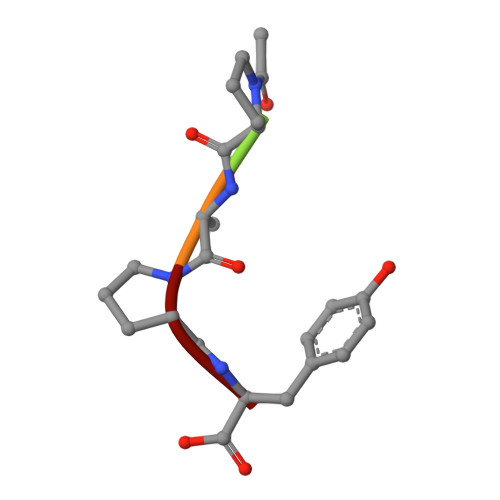
Structures of product and inhibitor complexes of Streptomyces griseus protease A at 1.8 A resolution. A model for serine protease catalysis.
James, M.N., Sielecki, A.R., Brayer, G.D., Delbaere, L.T., Bauer, C.A.(1980) J Mol Biology 144: 43-88
- PubMed: 6783761 Search on PubMed
- DOI: https://doi.org/10.1016/0022-2836(80)90214-4
- Primary Citation Related Structures:
3SGA, 4SGA, 5SGA