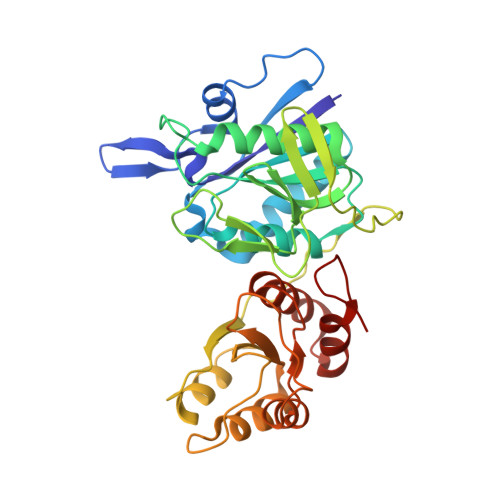
Structure and function of the thermostable L-asparaginase from Thermococcus kodakarensis.
Guo, J., Coker, A.R., Wood, S.P., Cooper, J.B., Chohan, S.M., Rashid, N., Akhtar, M.(2017) Acta Crystallogr D Struct Biol 73: 889-895
- PubMed: 29095161 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798317014711
- Primary Citation Related Structures:
5OT0 - PubMed Abstract:
L-Asparaginases catalyse the hydrolysis of asparagine to aspartic acid and ammonia. In addition, L-asparaginase is involved in the biosynthesis of amino acids such as lysine, methionine and threonine. These enzymes have been used as chemotherapeutic agents for the treatment of acute lymphoblastic leukaemia and other haematopoietic malignancies since the tumour cells cannot synthesize sufficient L-asparagine and are thus killed by deprivation of this amino acid. L-Asparaginases are also used in the food industry and have potential in the development of biosensors, for example for asparagine levels in leukaemia. The thermostable type I L-asparaginase from Thermococcus kodakarensis (TkA) is composed of 328 amino acids and forms homodimers in solution, with the highest catalytic activity being observed at pH 9.5 and 85°C. It has a K m value of 5.5 mM for L-asparagine, with no glutaminase activity being observed. The crystal structure of TkA has been determined at 2.18 Å resolution, confirming the presence of two α/β domains connected by a short linker region. The N-terminal domain contains a highly flexible β-hairpin which adopts `open' and `closed' conformations in different subunits of the solved TkA structure. In previously solved L-asparaginase structures this β-hairpin was only visible when in the `closed' conformation, whilst it is characterized with good electron density in all of the subunits of the TkA structure. A phosphate anion resides at the active site, which is formed by residues from both of the neighbouring monomers in the dimer. The high thermostability of TkA is attributed to the high arginine and salt-bridge content when compared with related mesophilic enzymes.
- Division of Medicine, University College London, Gower Street, London WC1E 6BT, England.
Organizational Affiliation: