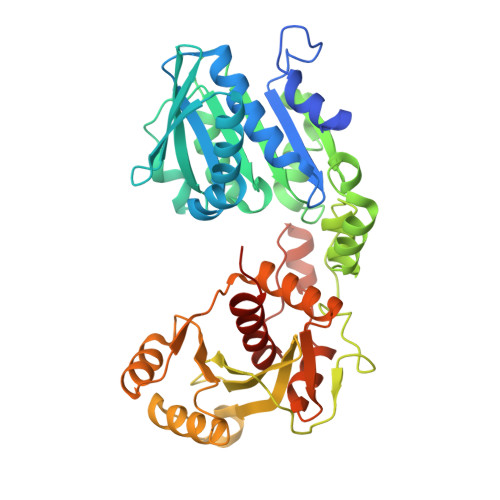
Structure of the large terminase from a hyperthermophilic virus reveals a unique mechanism for oligomerization and ATP hydrolysis.
Xu, R.G., Jenkins, H.T., Antson, A.A., Greive, S.J.(2017) Nucleic Acids Res 45: 13029-13042
- PubMed: 29069443
- DOI: https://doi.org/10.1093/nar/gkx947
- Primary Citation Related Structures:
5OE8, 5OE9, 5OEA, 5OEB, 5OEE - PubMed Abstract:
The crystal structure of the large terminase from the Geobacillus stearothermophilus bacteriophage D6E shows a unique relative orientation of the N-terminal adenosine triphosphatase (ATPase) and C-terminal nuclease domains. This monomeric 'initiation' state with the two domains 'locked' together is stabilized via a conserved C-terminal arm, which may interact with the portal protein during motor assembly, as predicted for several bacteriophages. Further work supports the formation of an active oligomeric state: (i) AUC data demonstrate the presence of oligomers; (ii) mutational analysis reveals a trans-arginine finger, R158, indispensable for ATP hydrolysis; (iii) the location of this arginine is conserved with the HerA/FtsK ATPase superfamily; (iv) a molecular docking model of the pentamer is compatible with the location of the identified arginine finger. However, this pentameric model is structurally incompatible with the monomeric 'initiation' state and is supported by the observed increase in kcat of ATP hydrolysis, from 7.8 ± 0.1 min-1 to 457.7 ± 9.2 min-1 upon removal of the C-terminal nuclease domain. Taken together, these structural, biophysical and biochemical data suggest a model where transition from the 'initiation' state into a catalytically competent pentameric state, is accompanied by substantial domain rearrangements, triggered by the removal of the C-terminal arm from the ATPase active site.
- York Structural Biology Laboratory, Department of Chemistry, University of York, York YO10 5DD, UK.
Organizational Affiliation: