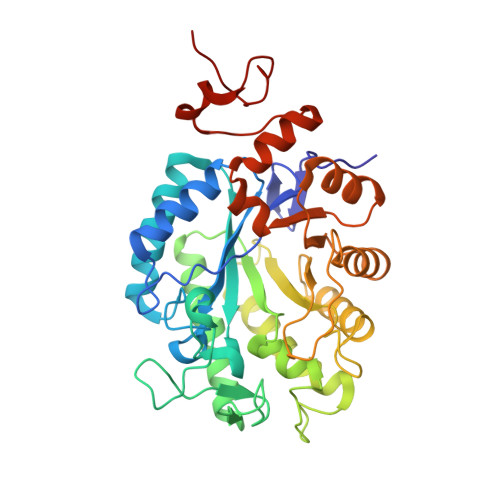
Structural investigation into the C-terminal extension of the ene-reductase from Ralstonia (Cupriavidus) metallidurans.
Opperman, D.J.(2017) Proteins 85: 2252-2257
- PubMed: 28833623 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25372
- Primary Citation Related Structures:
5OCS - PubMed Abstract:
Ene-reductases (ERs), or Old Yellow Enzymes, catalyze the asymmetric reduction of various activated alkenes. This class of biocatalysts is considered an attractive alternative to current chemical technologies for hydrogenation due to their high selectivity and specificity. Here the X-ray crystal structure of RmER, a "thermophilic"-like ER from Ralstonia (Cupriavidus) metallidurans, is reported. Unlike other members of this class of ERs, RmER is monomeric in solution which we previously related to its atypical elongated C-terminus. A typical dimer interface was however observed in our crystal structure, with the conserved Arg-"finger" forming part of the adjacent monomer's active site and the elongated C-terminus extending into the active site through contacting the "capping" domain. This dimerization also resulted in the loss of one FMN cofactor from each dimer pair. This potential transient dimerization and dissociation of FMN could conceivably explain the rapid rates previously observed when an FMN light-driven cofactor regeneration system was used during catalysis with RmER.
- Department of Biotechnology, University of the Free State, 205 Nelson Mandela Drive, Bloemfontein, 9300, South Africa.
Organizational Affiliation: