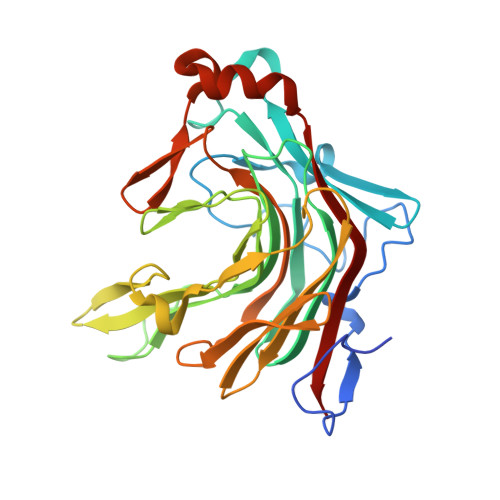
Structural insights into marine carbohydrate degradation by family GH16 kappa-carrageenases.
Matard-Mann, M., Bernard, T., Leroux, C., Barbeyron, T., Larocque, R., Prechoux, A., Jeudy, A., Jam, M., Nyvall Collen, P., Michel, G., Czjzek, M.(2017) J Biological Chem 292: 19919-19934
- PubMed: 29030427
- DOI: https://doi.org/10.1074/jbc.M117.808279
- Primary Citation Related Structures:
5OCQ, 5OCR - PubMed Abstract:
Carrageenans are sulfated α-1,3-β-1,4-galactans found in the cell wall of some red algae that are practically valuable for their gelation and biomimetic properties but also serve as a potential carbon source for marine bacteria. Carbohydrate degradation has been studied extensively for terrestrial plant/bacterial systems, but sulfation is not present in these cases, meaning the marine enzymes used to degrade carrageenans must possess unique features to recognize these modifications. To gain insights into these features, we have focused on κ-carrageenases from two distant bacterial phyla, which belong to glycoside hydrolase family 16 and cleave the β-1,4 linkage of κ-carrageenan. We have solved the crystal structure of the catalytic module of Zg CgkA from Zobellia galactanivorans at 1.66 Å resolution and compared it with the only other structure available, that of Pc CgkA from Pseudoalteromonas carrageenovora 9 T (ATCC 43555 T ). We also describe the first substrate complex in the inactivated mutant form of Pc CgkA at 1.7 Å resolution. The structural and biochemical comparison of these enzymes suggests key determinants that underlie the functional properties of this subfamily. In particular, we identified several arginine residues that interact with the polyanionic substrate, and confirmed the functional relevance of these amino acids using a targeted mutagenesis strategy. These results give new insight into the diversity of the κ-carrageenase subfamily. The phylogenetic analyses show the presence of several distinct clades of enzymes that relate to differences in modes of action or subtle differences within the same substrate specificity, matching the hybrid character of the κ-carrageenan polymer.
- From the Sorbonne Universités, UPMC Université Paris 06, CNRS, UMR 8227, Integrative Biology of Marine Models, Station Biologique de Roscoff, CS 90074 Roscoff, Bretagne, France.
Organizational Affiliation: