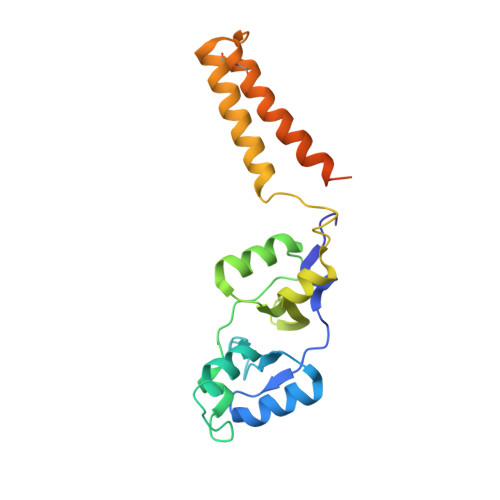
Structural and functional insights into the unique CBS-CP12 fusion protein family in cyanobacteria.
Hackenberg, C., Hakanpaa, J., Cai, F., Antonyuk, S., Eigner, C., Meissner, S., Laitaoja, M., Janis, J., Kerfeld, C.A., Dittmann, E., Lamzin, V.S.(2018) Proc Natl Acad Sci U S A 115: 7141-7146
- PubMed: 29915055 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1806668115
- Primary Citation Related Structures:
5NMU, 5NPL, 5NVD - PubMed Abstract:
Cyanobacteria are important photosynthetic organisms inhabiting a range of dynamic environments. This phylum is distinctive among photosynthetic organisms in containing genes encoding uncharacterized cystathionine β-synthase (CBS)-chloroplast protein (CP12) fusion proteins. These consist of two domains, each recognized as stand-alone photosynthetic regulators with different functions described in cyanobacteria (CP12) and plants (CP12 and CBSX). Here we show that CBS-CP12 fusion proteins are encoded in distinct gene neighborhoods, several unrelated to photosynthesis. Most frequently, CBS-CP12 genes are in a gene cluster with thioredoxin A (TrxA), which is prevalent in bloom-forming, marine symbiotic, and benthic mat cyanobacteria. Focusing on a CBS-CP12 from Microcystis aeruginosa PCC 7806 encoded in a gene cluster with TrxA, we reveal that the domain fusion led to the formation of a hexameric protein. We show that the CP12 domain is essential for hexamerization and contains an ordered, previously structurally uncharacterized N-terminal region. We provide evidence that CBS-CP12, while combining properties of both regulatory domains, behaves different from CP12 and plant CBSX. It does not form a ternary complex with phosphoribulokinase (PRK) and glyceraldehyde-3-phosphate dehydrogenase. Instead, CBS-CP12 decreases the activity of PRK in an AMP-dependent manner. We propose that the novel domain architecture and oligomeric state of CBS-CP12 expand its regulatory function beyond those of CP12 in cyanobacteria.
- European Molecular Biology Laboratory (EMBL), Deutsches Elektronen-Synchrotron (DESY), 22607 Hamburg, Germany; hackenberg@embl-hamburg.de victor@embl-hamburg.de.
Organizational Affiliation: