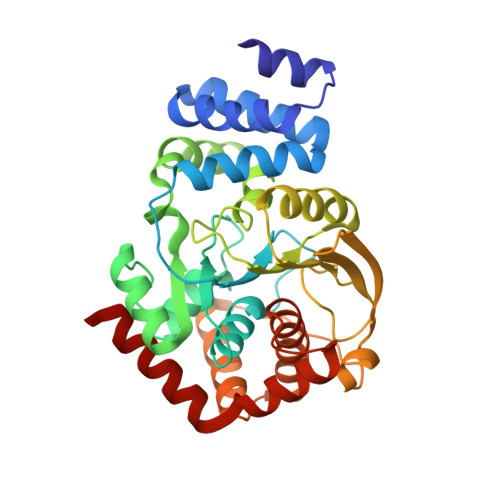
Anthranilate phosphoribosyltransferase from the hyperthermophilic archaeon Thermococcus kodakarensis shows maximum activity with zinc and forms a unique dimeric structure.
Perveen, S., Rashid, N., Tang, X.F., Imanaka, T., Papageorgiou, A.C.(2017) FEBS Open Bio 7: 1217-1230
- PubMed: 28781961
- DOI: https://doi.org/10.1002/2211-5463.12264
- Primary Citation Related Structures:
5NOE, 5NOF - PubMed Abstract:
Anthranilate phosphoribosyltransferase (TrpD) is involved in tryptophan biosynthesis, catalyzing the transfer of a phosphoribosyl group to anthranilate, leading to the generation of phosphoribosyl anthranilate. TrpD belongs to the phosphoribosyltransferase (PRT) superfamily and is the only member of the structural class IV. X-ray structures of TrpD from seven species have been solved to date. Here, functional and structural characterization of a recombinant TrpD from hyperthermophilic archaeon Thermococcus kodakarensis KOD1 ( Tk TrpD) was carried out. Contrary to previously characterized Mg 2+ -dependent TrpD enzymes, Tk TrpD was found to have a unique divalent cation dependency characterized by maximum activity in the presence of Zn 2+ (1580 μmol·min -1 ·mg -1 , the highest reported for any TrpD) followed by Ca 2+ (948 μmol·min -1 ·mg -1 ) and Mg 2+ (711 μmol·min -1 ·mg -1 ). Tk TrpD displayed an unusually low thermostability compared to other previously characterized proteins from T. kodakarensis KOD1. The crystal structure of Tk TrpD was determined in free form and in the presence of Zn 2+ to 1.9 and 2.4 Å resolutions, respectively. Tk TrpD structure displayed the typical PRT fold similar to other class IV PRTs, with a small N-terminal α-helical domain and a larger C-terminal α/β domain. Electron densities for Zn 2+ were identified at the expected zinc-binding motif, DE(217-218), of the enzyme in each subunit of the dimer. Two additional Zn 2+ were found at a new dimer interface formed in the presence of Zn 2+ . A fifth Zn 2+ was found bound to Glu118 at crystal lattice contacts and a sixth one was ligated with Glu235. Based on the Tk TrpD-Zn 2+ structure, it is suggested that the formation of a new dimer may be responsible for the higher enzyme activity of Tk TrpD in the presence of Zn 2+ ions.
- School of Biological Sciences University of the Punjab Lahore Pakistan.
Organizational Affiliation: