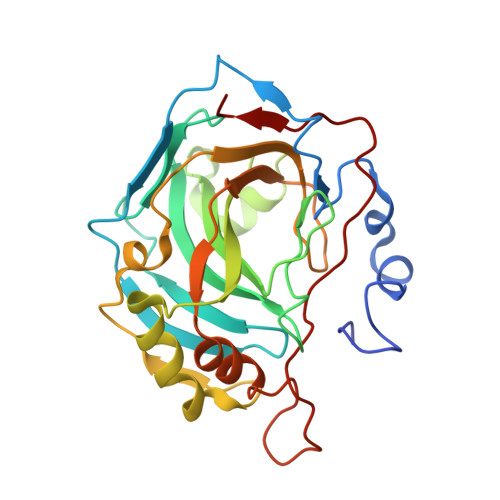
Probing Molecular Interactions between Human Carbonic Anhydrases (hCAs) and a Novel Class of Benzenesulfonamides.
Bruno, E., Buemi, M.R., Di Fiore, A., De Luca, L., Ferro, S., Angeli, A., Cirilli, R., Sadutto, D., Alterio, V., Monti, S.M., Supuran, C.T., De Simone, G., Gitto, R.(2017) J Med Chem 60: 4316-4326
- PubMed: 28453941 Search on PubMed
- DOI: https://doi.org/10.1021/acs.jmedchem.7b00264
- Primary Citation Related Structures:
5N0D, 5N0E - PubMed Abstract:
On the basis of X-ray crystallographic studies of the complex of hCA II with 4-(3,4-dihydro-1H-isoquinoline-2-carbonyl)benzenesulfonamide (3) (PDB code 4Z1J ), a novel series of 4-(1-aryl-3,4-dihydro-1H-isoquinolin-2-carbonyl)benzenesulfonamides (23-33) was designed. Specifically, our idea was to improve the selectivity toward druggable isoforms through the introduction of additional hydrophobic/hydrophilic functionalities. Among the synthesized and tested compounds, the (R,S)-4-(6,7-dihydroxy-1-phenyl-3,4-tetrahydroisoquinoline-1H-2-carbonyl)benzenesulfonamide (30) exhibited a remarkable inhibition for the brain-expressed hCA VII (K i = 0.20 nM) and selectivity over wider distributed hCA I and hCA II isoforms. By enantioselective HPLC, we solved the racemic mixture and ascertained that the two enantiomers (30a and 30b) are equiactive inhibitors for hCA VII. Crystallographic and docking studies revealed the main interactions of these inhibitors into the carbonic anhydrase (CA) catalytic site, thus highlighting the relevant role of nonpolar contacts for this class of hCA inhibitors.
- Dipartimento di Scienze Chimiche, Biologiche, Farmaceutiche ed Ambientali (CHIBIOFARAM), Università degli Studi di Messina , Viale Annunziata, I-98168 Messina, Italy.
Organizational Affiliation: