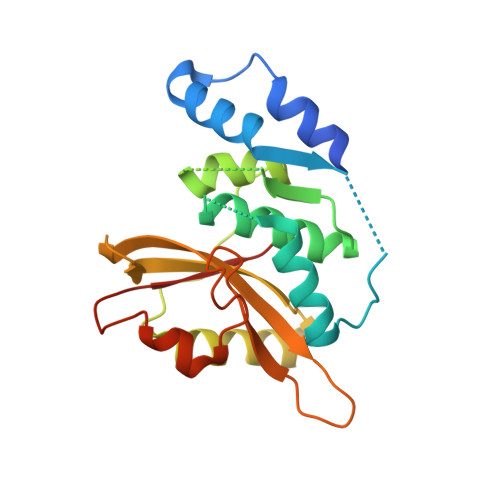
The Salmonella Effector SpvD Is a Cysteine Hydrolase with a Serovar-specific Polymorphism Influencing Catalytic Activity, Suppression of Immune Responses, and Bacterial Virulence.
Grabe, G.J., Zhang, Y., Przydacz, M., Rolhion, N., Yang, Y., Pruneda, J.N., Komander, D., Holden, D.W., Hare, S.A.(2016) J Biological Chem 291: 25853-25863
- PubMed: 27789710
- DOI: https://doi.org/10.1074/jbc.M116.752782
- Primary Citation Related Structures:
5LQ6, 5LQ7 - PubMed Abstract:
Many bacterial pathogens secrete virulence (effector) proteins that interfere with immune signaling in their host. SpvD is a Salmonella enterica effector protein that we previously demonstrated to negatively regulate the NF-κB signaling pathway and promote virulence of S. enterica serovar Typhimurium in mice. To shed light on the mechanistic basis for these observations, we determined the crystal structure of SpvD and show that it adopts a papain-like fold with a characteristic cysteine-histidine-aspartate catalytic triad comprising Cys-73, His-162, and Asp-182. SpvD possessed an in vitro deconjugative activity on aminoluciferin-linked peptide and protein substrates in vitro A C73A mutation abolished SpvD activity, demonstrating that an intact catalytic triad is required for its function. Taken together, these results strongly suggest that SpvD is a cysteine protease. The amino acid sequence of SpvD is highly conserved across different S. enterica serovars, but residue 161, located close to the catalytic triad, is variable, with serovar Typhimurium SpvD having an arginine and serovar Enteritidis a glycine at this position. This variation affected hydrolytic activity of the enzyme on artificial substrates and can be explained by substrate accessibility to the active site. Interestingly, the SpvD G161 variant more potently inhibited NF-κB-mediated immune responses in cells in vitro and increased virulence of serovar Typhimurium in mice. In summary, our results explain the biochemical basis for the effect of virulence protein SpvD and demonstrate that a single amino acid polymorphism can affect the overall virulence of a bacterial pathogen in its host.
- From the Section of Microbiology and.
Organizational Affiliation: