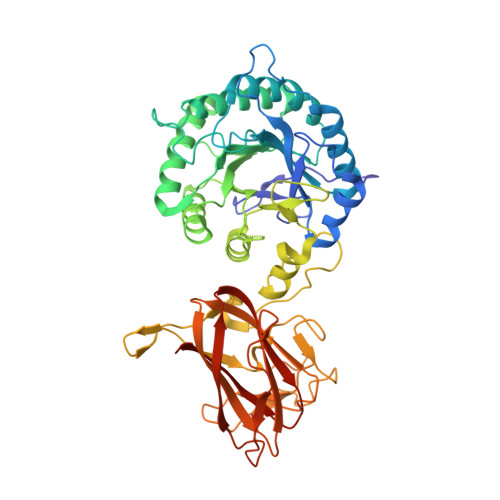
The Mechanism by Which Arabinoxylanases Can Recognize Highly Decorated Xylans.
Labourel, A., Crouch, L.I., Bras, J.L., Jackson, A., Rogowski, A., Gray, J., Yadav, M.P., Henrissat, B., Fontes, C.M., Gilbert, H.J., Najmudin, S., Basle, A., Cuskin, F.(2016) J Biological Chem 291: 22149-22159
- PubMed: 27531750 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1074/jbc.M116.743948
- Primary Citation Related Structures:
5G56, 5LA0, 5LA1, 5LA2 - PubMed Abstract:
The enzymatic degradation of plant cell walls is an important biological process of increasing environmental and industrial significance. Xylan, a major component of the plant cell wall, consists of a backbone of β-1,4-xylose (Xylp) units that are often decorated with arabinofuranose (Araf) side chains. A large penta-modular enzyme, CtXyl5A, was shown previously to specifically target arabinoxylans. The mechanism of substrate recognition displayed by the enzyme, however, remains unclear. Here we report the crystal structure of the arabinoxylanase and the enzyme in complex with ligands. The data showed that four of the protein modules adopt a rigid structure, which stabilizes the catalytic domain. The C-terminal non-catalytic carbohydrate binding module could not be observed in the crystal structure, suggesting positional flexibility. The structure of the enzyme in complex with Xylp-β-1,4-Xylp-β-1,4-Xylp-[α-1,3-Araf]-β-1,4-Xylp showed that the Araf decoration linked O 3 to the xylose in the active site is located in the pocket (-2* subsite) that abuts onto the catalytic center. The -2* subsite can also bind to Xylp and Arap, explaining why the enzyme can utilize xylose and arabinose as specificity determinants. Alanine substitution of Glu 68 , Tyr 92 , or Asn 139 , which interact with arabinose and xylose side chains at the -2* subsite, abrogates catalytic activity. Distal to the active site, the xylan backbone makes limited apolar contacts with the enzyme, and the hydroxyls are solvent-exposed. This explains why CtXyl5A is capable of hydrolyzing xylans that are extensively decorated and that are recalcitrant to classic endo-xylanase attack.
- From the Institute for Cell and Molecular Biosciences, Newcastle University, Newcastle upon Tyne NE2 4HH, United Kingdom.
Organizational Affiliation: