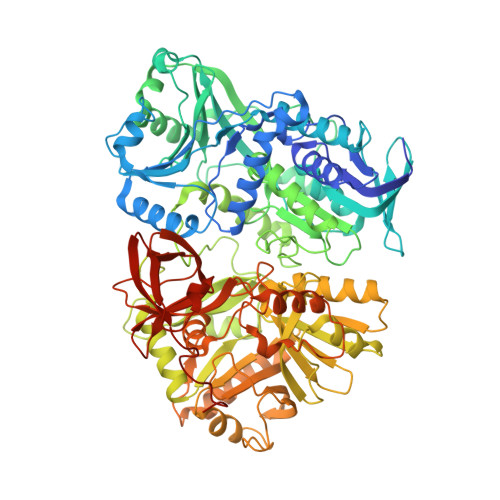
Structure and biochemical properties of recombinant human dimethylglycine dehydrogenase and comparison to the disease-related H109R variant.
Augustin, P., Hromic, A., Pavkov-Keller, T., Gruber, K., Macheroux, P.(2016) FEBS J 283: 3587-3603
- PubMed: 27486859 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1111/febs.13828
- Primary Citation Related Structures:
5L46 - PubMed Abstract:
The human dimethylglycine dehydrogenase (hDMGDH) is a flavin adenine dinucleotide (FAD)- and tetrahydrofolate (THF)-dependent, mitochondrial matrix enzyme taking part in choline degradation, one-carbon metabolism and electron transfer to the respiratory chain. The rare natural variant H109R causes dimethylglycine dehydrogenase deficiency leading to increased blood and urinary dimethylglycine concentrations. A detailed biochemical and structural characterization of hDMGDH was thus far hampered by insufficient heterologous expression of the protein. In the present study, we report the development of an intracellular, heterologous expression system in Komagataella phaffii (formerly known as Pichia pastoris) providing the opportunity to determine kinetic parameters, spectroscopic properties, thermostability, and the redox potential of hDMGDH. Moreover, we have successfully crystallized the wild-type enzyme and determined the structure to 3.1-Å resolution. The structure-based analysis of our biochemical data provided new insights into the kinetic properties of the enzyme in particular with respect to oxygen reactivity. A comparative study with the H109R variant demonstrated that the variant suffers from decreased protein stability, cofactor saturation, and substrate affinity. Structural data are available in the PDB database under the accession number 5L46.
- Institute of Biochemistry, Graz University of Technology, Austria.
Organizational Affiliation: