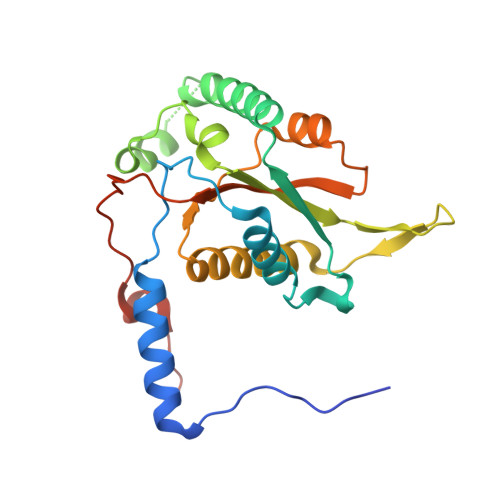
Active Site Binding Is Not Sufficient for Reductive Deiodination by Iodotyrosine Deiodinase.
Ingavat, N., Kavran, J.M., Sun, Z., Rokita, S.E.(2017) Biochemistry 56: 1130-1139
- PubMed: 28157283 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acs.biochem.6b01308
- Primary Citation Related Structures:
5KO7, 5KO8, 5KRD - PubMed Abstract:
The minimal requirements for substrate recognition and turnover by iodotyrosine deiodinase were examined to learn the basis for its catalytic specificity. This enzyme is crucial for iodide homeostasis and the generation of thyroid hormone in chordates. 2-Iodophenol binds only very weakly to the human enzyme and is dehalogenated with a k cat /K m that is more than 4 orders of magnitude lower than that for iodotyrosine. This discrimination likely protects against a futile cycle of iodinating and deiodinating precursors of thyroid hormone biosynthesis. Surprisingly, a very similar catalytic selectivity was expressed by a bacterial homologue from Haliscomenobacter hydrossis. In this example, discrimination was not based on affinity since 4-cyano-2-iodophenol bound to the bacterial deiodinase with a K d lower than that of iodotyrosine and yet was not detectably deiodinated. Other phenols including 2-iodophenol were deiodinated but only very inefficiently. Crystal structures of the bacterial enzyme with and without bound iodotyrosine are nearly superimposable and quite similar to the corresponding structures of the human enzyme. Likewise, the bacterial enzyme is activated for single electron transfer after binding to the substrate analogue fluorotyrosine as previously observed with the human enzyme. A cocrystal structure of bacterial deiodinase and 2-iodophenol indicates that this ligand stacks on the active site flavin mononucleotide (FMN) in a orientation analogous to that of bound iodotyrosine. However, 2-iodophenol association is not sufficient to activate the FMN chemistry required for catalysis, and thus the bacterial enzyme appears to share a similar specificity for halotyrosines even though their physiological roles are likely very different from those in humans.
- Department of Chemistry, Johns Hopkins University , 3400 North Charles Street, Baltimore, Maryland 21218, United States.
Organizational Affiliation: