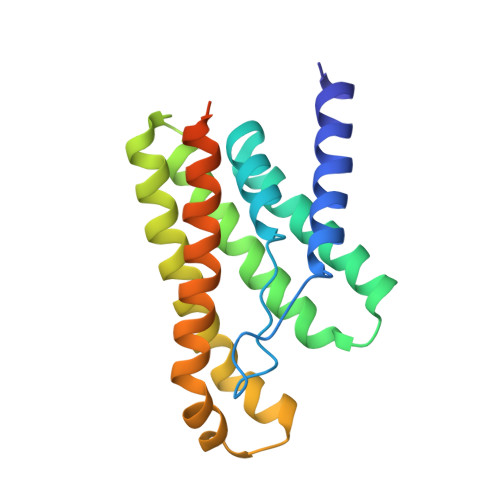
An Aromatic Cap Seals the Substrate Binding Site in an ECF-Type S Subunit for Riboflavin.
Karpowich, N.K., Song, J., Wang, D.N.(2016) J Mol Biology 428: 3118-3130
- PubMed: 27312125 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.jmb.2016.06.003
- Primary Citation Related Structures:
5KBW, 5KC0, 5KC4 - PubMed Abstract:
ECF transporters are a family of active membrane transporters for essential micronutrients, such as vitamins and trace metals. Found exclusively in archaea and bacteria, these transporters are composed of four subunits: an integral membrane substrate-binding subunit (EcfS), a transmembrane coupling subunit (EcfT), and two ATP-binding cassette ATPases (EcfA and EcfA'). We have characterized the structural basis of substrate binding by the EcfS subunit for riboflavin from Thermotoga maritima, TmRibU. TmRibU binds riboflavin with high affinity, and the protein-substrate complex is exceptionally stable in solution. The crystal structure of riboflavin-bound TmRibU reveals an electronegative binding pocket at the extracellular surface in which the substrate is completely buried. Analysis of the intermolecular contacts indicates that nearly every available substrate hydrogen bond is satisfied. A conserved aromatic residue at the extracellular end of TM5, Tyr130, caps the binding site to generate a substrate-bound, occluded state, and non-conservative mutation of Tyr130 reduces the stability of this conformation. Using a novel fluorescence binding assay, we find that an aromatic residue at this position is essential for high-affinity substrate binding. Comparison with other S subunit structures suggests that TM5 and Loop5-6 contain a dynamic, conserved motif that plays a key role in gating substrate entry and release by S subunits of ECF transporters.
- The Helen L. and Martin S. Kimmel Center for Biology and Medicine at the Skirball Institute of Biomolecular Medicine, and Department of Cell Biology, New York University School of Medicine, 540 First Avenue, New York, NY 10016, USA. Electronic address: Nathan.Karpowich@med.nyu.edu.
Organizational Affiliation: