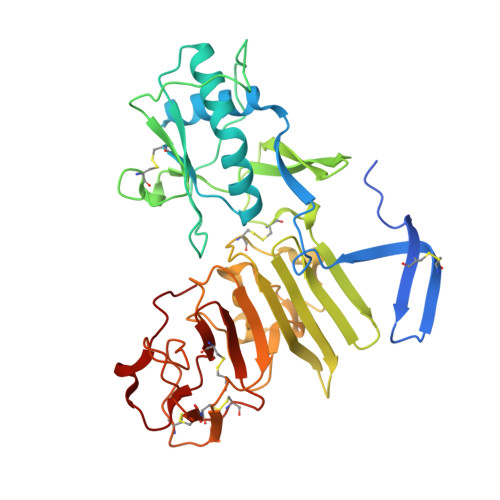
Extended surface for membrane association in Zika virus NS1 structure.
Brown, W.C., Akey, D.L., Konwerski, J.R., Tarrasch, J.T., Skiniotis, G., Kuhn, R.J., Smith, J.L.(2016) Nat Struct Mol Biol 23: 865-867
- PubMed: 27455458 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/nsmb.3268
- Primary Citation Related Structures:
5K6K - PubMed Abstract:
The Zika virus, which has been implicated in an increase in neonatal microcephaly and Guillain-Barré syndrome, has spread rapidly through tropical regions of the world. The virulence protein NS1 functions in genome replication and host immune-system modulation. Here, we report the crystal structure of full-length Zika virus NS1, revealing an elongated hydrophobic surface for membrane association and a polar surface that varies substantially among flaviviruses.
- Life Sciences Institute, University of Michigan, Ann Arbor, Michigan, USA.
Organizational Affiliation: