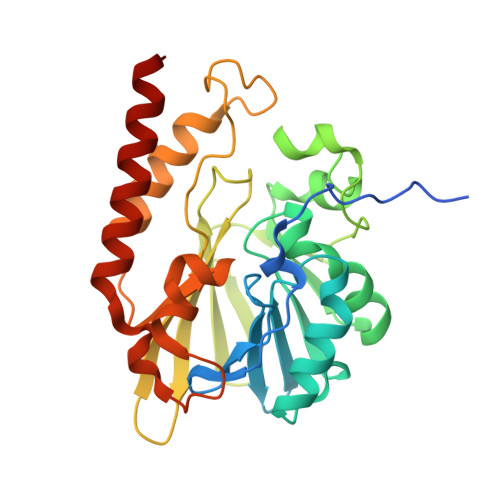
Crystal Structure of the Metallo-beta-Lactamase GOB in the Periplasmic Dizinc Form Reveals an Unusual Metal Site.
Moran-Barrio, J., Lisa, M.N., Larrieux, N., Drusin, S.I., Viale, A.M., Moreno, D.M., Buschiazzo, A., Vila, A.J.(2016) Antimicrob Agents Chemother 60: 6013-6022
- PubMed: 27458232
- DOI: https://doi.org/10.1128/AAC.01067-16
- Primary Citation of Related Structures:
5K0W - PubMed Abstract:
Metallo-beta-lactamases (MBLs) are broad-spectrum, Zn(II)-dependent lactamases able to confer resistance to virtually every β-lactam antibiotic currently available. The large diversity of active-site structures and metal content among MBLs from different sources has limited the design of a pan-MBL inhibitor. GOB-18 is a divergent MBL from subclass B3 that is expressed by the opportunistic Gram-negative pathogen Elizabethkingia meningoseptica This MBL is atypical, since several residues conserved in B3 enzymes (such as a metal ligand His) are substituted in GOB enzymes. Here, we report the crystal structure of the periplasmic di-Zn(II) form of GOB-18. This enzyme displays a unique active-site structure, with residue Gln116 coordinating the Zn1 ion through its terminal amide moiety, replacing a ubiquitous His residue. This situation contrasts with that of B2 MBLs, where an equivalent His116Asn substitution leads to a di-Zn(II) inactive species. Instead, both the mono- and di-Zn(II) forms of GOB-18 are active against penicillins, cephalosporins, and carbapenems. In silico docking and molecular dynamics simulations indicate that residue Met221 is not involved in substrate binding, in contrast to Ser221, which otherwise is conserved in most B3 enzymes. These distinctive features are conserved in recently reported GOB orthologues in environmental bacteria. These findings provide valuable information for inhibitor design and also posit that GOB enzymes have alternative functions.
- Departamento de Química Biológica and Instituto de Biología Molecular y Celular de Rosario (IBR, CONICET-UNR), Facultad de Ciencias Bioquímicas y Farmacéuticas, Universidad Nacional de Rosario, Rosario, Argentina.
Organizational Affiliation: