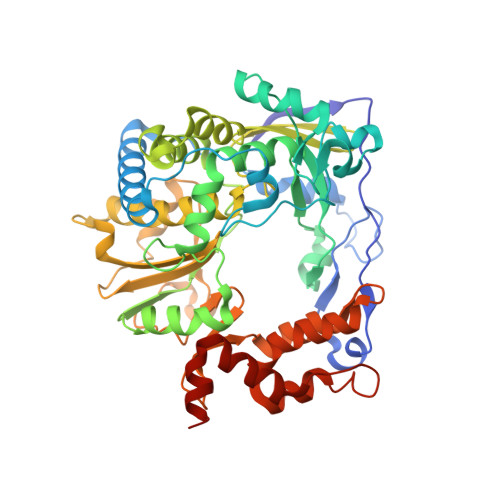
Both cis and trans Activities of Foot-and-Mouth Disease Virus 3D Polymerase Are Essential for Viral RNA Replication.
Herod, M.R., Ferrer-Orta, C., Loundras, E.A., Ward, J.C., Verdaguer, N., Rowlands, D.J., Stonehouse, N.J.(2016) J Virol 90: 6864-6883
- PubMed: 27194768 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JVI.00469-16
- Primary Citation Related Structures:
5JXS - PubMed Abstract:
The Picornaviridae is a large family of positive-sense RNA viruses that contains numerous human and animal pathogens, including foot-and-mouth disease virus (FMDV). The picornavirus replication complex comprises a coordinated network of protein-protein and protein-RNA interactions involving multiple viral and host-cellular factors. Many of the proteins within the complex possess multiple roles in viral RNA replication, some of which can be provided in trans (i.e., via expression from a separate RNA molecule), while others are required in cis (i.e., expressed from the template RNA molecule). In vitro studies have suggested that multiple copies of the RNA-dependent RNA polymerase (RdRp) 3D are involved in the viral replication complex. However, it is not clear whether all these molecules are catalytically active or what other function(s) they provide. In this study, we aimed to distinguish between catalytically active 3D molecules and those that build a replication complex. We report a novel nonenzymatic cis-acting function of 3D that is essential for viral-genome replication. Using an FMDV replicon in complementation experiments, our data demonstrate that this cis-acting role of 3D is distinct from the catalytic activity, which is predominantly trans acting. Immunofluorescence studies suggest that both cis- and trans-acting 3D molecules localize to the same cellular compartment. However, our genetic and structural data suggest that 3D interacts in cis with RNA stem-loops that are essential for viral RNA replication. This study identifies a previously undescribed aspect of picornavirus replication complex structure-function and an important methodology for probing such interactions further. Foot-and-mouth disease virus (FMDV) is an important animal pathogen responsible for foot-and-mouth disease. The disease is endemic in many parts of the world with outbreaks within livestock resulting in major economic losses. Propagation of the viral genome occurs within replication complexes, and understanding this process can facilitate the development of novel therapeutic strategies. Many of the nonstructural proteins involved in replication possess multiple functions in the viral life cycle, some of which can be supplied to the replication complex from a separate genome (i.e., in trans) while others must originate from the template (i.e., in cis). Here, we present an analysis of cis and trans activities of the RNA-dependent RNA polymerase 3D. We demonstrate a novel cis-acting role of 3D in replication. Our data suggest that this role is distinct from its enzymatic functions and requires interaction with the viral genome. Our data further the understanding of genome replication of this important pathogen.
- School of Molecular and Cellular Biology, Faculty of Biological Sciences and Astbury Centre for Structural Molecular Biology, University of Leeds, Leeds, United Kingdom.
Organizational Affiliation: