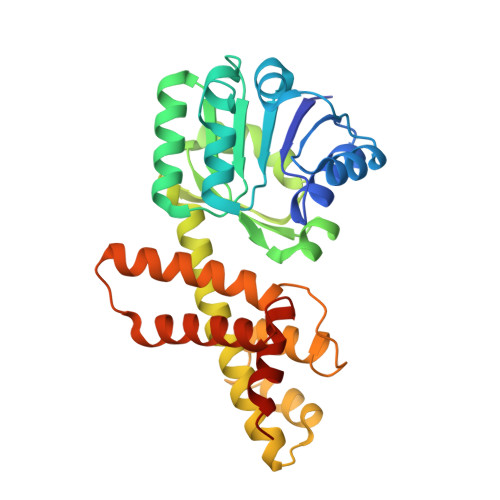
Structural and biochemical characterization of the Bacillus cereus 3-hydroxyisobutyrate dehydrogenase
Park, S.C., Kim, P.H., Lee, G.S., Kang, S.G., Ko, H.J., Yoon, S.I.(2016) Biochem Biophys Res Commun 474: 522-527
- PubMed: 27120461 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2016.04.126
- Primary Citation Related Structures:
5JE8 - PubMed Abstract:
The 3-hydroxyisobutyrate dehydrogenase (HIBADH) family catalyzes the NAD(+)- or NADP(+)-dependent oxidation of various β-hydroxyacid substrates into their cognate semialdehydes for diverse metabolic pathways. Because HIBADH group members exhibit different substrate specificities, the substrate-recognition mode of each enzyme should be individually characterized. In the current study, we report the biochemical and structural analysis of a HIBADH group enzyme from Bacillus cereus (bcHIBADH). bcHIBADH mediates a dehydrogenation reaction on S-3-hydroxyisobutyrate substrate with high catalytic efficiency in an NAD(+)-dependent manner; it also oxidizes l-serine and 3-hydroxypropionate with lower activity. bcHIBADH consists of two domains and is further assembled into a functional dimer rather than a tetramer that has been commonly observed in other prokaryotic HIBADH group members. In the bcHIBADH structure, the interdomain cleft forms a putative active site and simultaneously accommodates both an NAD(+) cofactor and a substrate mimic. Our structure-based comparative analysis highlights structural motifs that are important in the cofactor and substrate recognition of the HIBADH group.
- Division of Biomedical Convergence, College of Biomedical Science, Kangwon National University, Chuncheon 24341, Republic of Korea.
Organizational Affiliation: