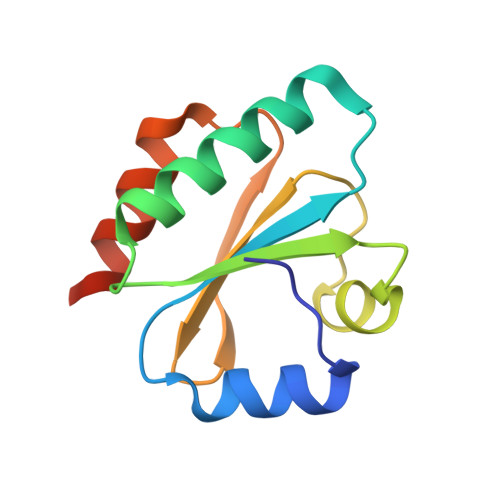
Computational Redesign of Thioredoxin Is Hypersensitive toward Minor Conformational Changes in the Backbone Template.
Johansson, K.E., Johansen, N.T., Christensen, S., Horowitz, S., Bardwell, J.C., Olsen, J.G., Willemoes, M., Lindorff-Larsen, K., Ferkinghoff-Borg, J., Hamelryck, T., Winther, J.R.(2016) J Mol Biology 428: 4361-4377
- PubMed: 27659562
- DOI: https://doi.org/10.1016/j.jmb.2016.09.013
- Primary Citation Related Structures:
5J7D - PubMed Abstract:
Despite the development of powerful computational tools, the full-sequence design of proteins still remains a challenging task. To investigate the limits and capabilities of computational tools, we conducted a study of the ability of the program Rosetta to predict sequences that recreate the authentic fold of thioredoxin. Focusing on the influence of conformational details in the template structures, we based our study on 8 experimentally determined template structures and generated 120 designs from each. For experimental evaluation, we chose six sequences from each of the eight templates by objective criteria. The 48 selected sequences were evaluated based on their progressive ability to (1) produce soluble protein in Escherichia coli and (2) yield stable monomeric protein, and (3) on the ability of the stable, soluble proteins to adopt the target fold. Of the 48 designs, we were able to synthesize 32, 20 of which resulted in soluble protein. Of these, only two were sufficiently stable to be purified. An X-ray crystal structure was solved for one of the designs, revealing a close resemblance to the target structure. We found a significant difference among the eight template structures to realize the above three criteria despite their high structural similarity. Thus, in order to improve the success rate of computational full-sequence design methods, we recommend that multiple template structures are used. Furthermore, this study shows that special care should be taken when optimizing the geometry of a structure prior to computational design when using a method that is based on rigid conformations.
- Linderstrøm-Lang Centre for Protein Science, Department of Biology, University of Copenhagen, Ole Maaløes Vej 5, Copenhagen DK-2200, Denmark.
Organizational Affiliation: