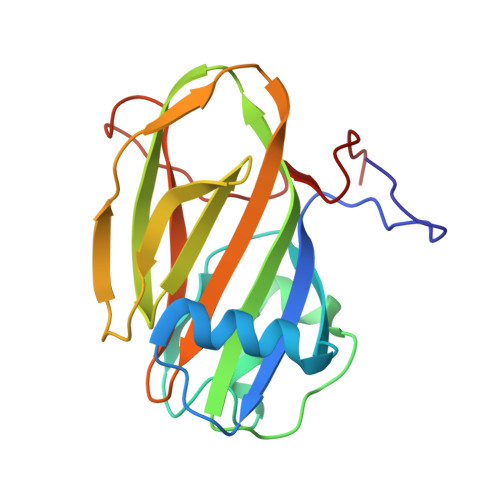
Structure of a 1.5-MDa adhesin that binds its Antarctic bacterium to diatoms and ice.
Guo, S., Stevens, C.A., Vance, T.D.R., Olijve, L.L.C., Graham, L.A., Campbell, R.L., Yazdi, S.R., Escobedo, C., Bar-Dolev, M., Yashunsky, V., Braslavsky, I., Langelaan, D.N., Smith, S.P., Allingham, J.S., Voets, I.K., Davies, P.L.(2017) Sci Adv 3: e1701440-e1701440
- PubMed: 28808685
- DOI: https://doi.org/10.1126/sciadv.1701440
- Primary Citation Related Structures:
5IRB, 5IX9, 5J6Y, 5JUH, 5K8G - PubMed Abstract:
Bacterial adhesins are modular cell-surface proteins that mediate adherence to other cells, surfaces, and ligands. The Antarctic bacterium Marinomonas primoryensis uses a 1.5-MDa adhesin comprising over 130 domains to position it on ice at the top of the water column for better access to oxygen and nutrients. We have reconstructed this 0.6-μm-long adhesin using a "dissect and build" structural biology approach and have established complementary roles for its five distinct regions. Domains in region I (RI) tether the adhesin to the type I secretion machinery in the periplasm of the bacterium and pass it through the outer membrane. RII comprises ~120 identical immunoglobulin-like β-sandwich domains that rigidify on binding Ca 2+ to project the adhesion regions RIII and RIV into the medium. RIII contains ligand-binding domains that join diatoms and bacteria together in a mixed-species community on the underside of sea ice where incident light is maximal. RIV is the ice-binding domain, and the terminal RV domain contains several "repeats-in-toxin" motifs and a noncleavable signal sequence that target proteins for export via the type I secretion system. Similar structural architecture is present in the adhesins of many pathogenic bacteria and provides a guide to finding and blocking binding domains to weaken infectivity.
- Protein Function Discovery Group and Department of Biomedical and Molecular Sciences, Queen's University, Kingston, Ontario K7L 3N6, Canada.
Organizational Affiliation: