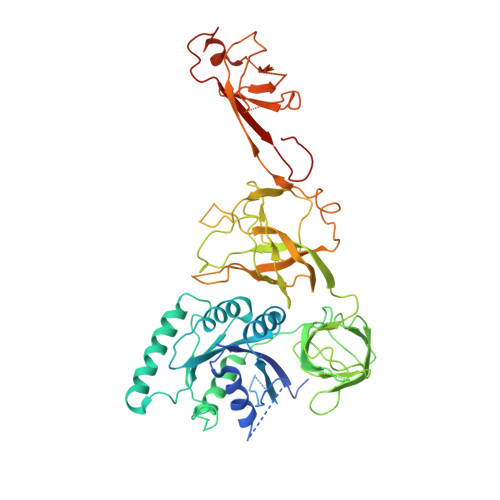
Crystal structures of the human elongation factor eEFSec suggest a non-canonical mechanism for selenocysteine incorporation.
Dobosz-Bartoszek, M., Pinkerton, M.H., Otwinowski, Z., Chakravarthy, S., Soll, D., Copeland, P.R., Simonovic, M.(2016) Nat Commun 7: 12941-12941
- PubMed: 27708257 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/ncomms12941
- Primary Citation Related Structures:
5IZK, 5IZL, 5IZM - PubMed Abstract:
Selenocysteine is the only proteinogenic amino acid encoded by a recoded in-frame UGA codon that does not operate as the canonical opal stop codon. A specialized translation elongation factor, eEFSec in eukaryotes and SelB in prokaryotes, promotes selenocysteine incorporation into selenoproteins by a still poorly understood mechanism. Our structural and biochemical results reveal that four domains of human eEFSec fold into a chalice-like structure that has similar binding affinities for GDP, GTP and other guanine nucleotides. Surprisingly, unlike in eEF1A and EF-Tu, the guanine nucleotide exchange does not cause a major conformational change in domain 1 of eEFSec, but instead induces a swing of domain 4. We propose that eEFSec employs a non-canonical mechanism involving the distinct C-terminal domain 4 for the release of the selenocysteinyl-tRNA during decoding on the ribosome.
- Department of Biochemistry and Molecular Genetics, University of Illinois at Chicago, Chicago, Illinois 60607, USA.
Organizational Affiliation: