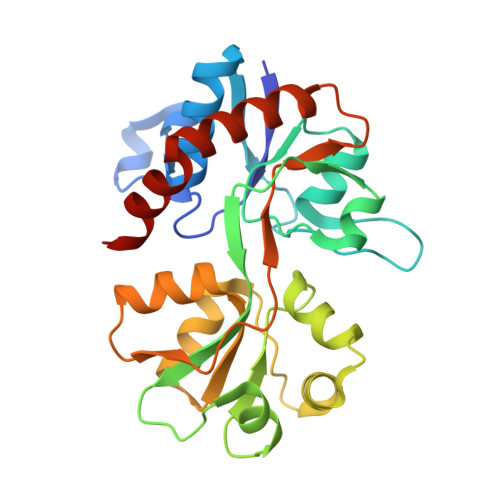
The Structure of a High-Affinity Kainate Receptor: GluK4 Ligand-Binding Domain Crystallized with Kainate.
Kristensen, O., Kristensen, L.B., Mollerud, S., Frydenvang, K., Pickering, D.S., Kastrup, J.S.(2016) Structure 24: 1582-1589
- PubMed: 27524200
- DOI: https://doi.org/10.1016/j.str.2016.06.019
- Primary Citation of Related Structures:
5IKB - PubMed Abstract:
Ionotropic glutamate receptors play a key role in fast neurotransmission in the CNS and have been linked to several neurological diseases and disorders. One subfamily is the kainate receptors, which are grouped into low-affinity (GluK1-3) and high-affinity (GluK4-5) receptors based on their affinity for kainate. Although structures of the ligand-binding domain (LBD) of all low-affinity kainate receptors have been reported, no structures of the high-affinity receptor subunits are available. Here, we present the X-ray structure of GluK4-LBD with kainate at 2.05 Å resolution, together with thermofluor and radiolabel binding affinity data. Whereas binding-site residues in GluK4 are most similar to the AMPA receptor subfamily, the domain closure and D1-D2 interlobe contacts induced by kainate are similar to the low-affinity kainate receptor GluK1. These observations provide a likely explanation for the high binding affinity of kainate at GluK4-LBD.
- Department of Drug Design and Pharmacology, Faculty of Health and Medical Sciences, University of Copenhagen, Jagtvej 162, 2100 Copenhagen Ø, Denmark.
Organizational Affiliation: