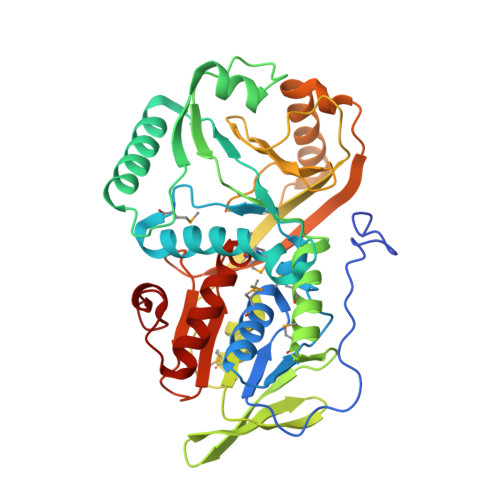
Crystal structure of a membrane-bound l-amino acid deaminase from Proteus vulgaris
Ju, Y., Tong, S., Gao, Y., Zhao, W., Liu, Q., Gu, Q., Xu, J., Niu, L., Teng, M., Zhou, H.(2016) J Struct Biol 195: 306-315
- PubMed: 27422658 Search on PubMed
- DOI: https://doi.org/10.1016/j.jsb.2016.07.008
- Primary Citation Related Structures:
5HXW, 5I39 - PubMed Abstract:
l-amino acid oxidases/deaminases (LAAOs/LAADs) are a class of oxidoreductases catalyzing the oxidative deamination of l-amino acids to α-keto acids. They are widely distributed in eukaryotic and prokaryotic organisms, and exhibit diverse substrate specificity, post-translational modifications and cellular localization. While LAAOs isolated from snake venom have been extensively characterized, the structures and functions of LAAOs from other species are largely unknown. Here, we reported crystal structure of a bacterial membrane-bound LAAD from Proteus vulgaris (pvLAAD) in complex with flavin adenine dinucleotide (FAD). We found that the overall fold of pvLAAD does not resemble typical LAAOs. Instead it, is similar to d-amino acid oxidases (DAAOs) with an additional hydrophobic insertion module on protein surface. Structural analysis and liposome-binding assays suggested that the hydrophobic module serves as an extra membrane-binding site for LAADs. Bacteria from genera Proteus and Providencia were found to encode two classes of membrane-bound LAADs. Based on our structure, the key roles of residues Q278 and L317 in substrate selectivity were proposed and biochemically analyzed. While LAADs on the membrane were proposed to transfer electrons to respiratory chain for FAD re-oxidization, we observed that the purified pvLAAD could generate a significant amount of hydrogen peroxide in vitro, suggesting it could use dioxygen to directly re-oxidize FADH2 as what typical LAAOs usually do. These findings provide a novel insights for a better understanding this class of enzymes and will help developing biocatalysts for industrial applications.
- Research Center for Structural Biology and Drug Discovery, School of Pharmaceutical Sciences, Sun Yat-sen University, Guangzhou 510006, China.
Organizational Affiliation: