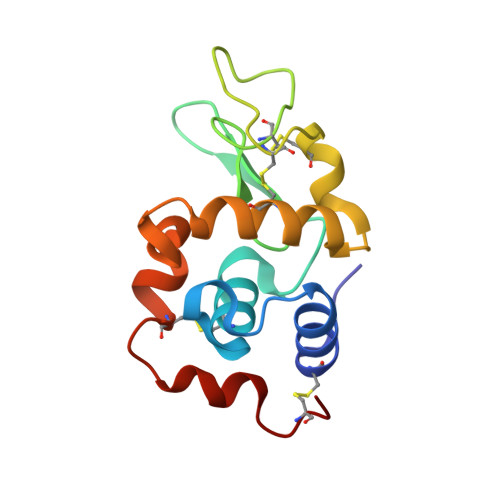
Re-refinement of 4g4a: room-temperature X-ray diffraction study of cisplatin and its binding to His15 of HEWL after 14 months chemical exposure in the presence of DMSO.
Tanley, S.W., Schreurs, A.M., Kroon-Batenburg, L.M., Helliwell, J.R.(2016) Acta Crystallogr F Struct Biol Commun 72: 253-254
- PubMed: 26948967 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X16000856
- Primary Citation Related Structures:
5HLL - PubMed Abstract:
A re-refinement of 4g4a, the room-temperature X-ray diffraction study of cisplatin and its binding to His15 of HEWL after 14 months chemical exposure in the presence of DMSO is published as an addendum to Tanley et al. [(2012), Acta Cryst. F68, 1300-1306]. This example illustrates the benefits of sharing raw diffraction images, as well as structure factors and molecular coordinates, as the diffraction resolution of the study is now much improved at 1.70 Å.
- School of Chemistry, Faculty of Engineering and Physical Sciences, University of Manchester, Brunswick Street, Manchester M13 9PL, England.
Organizational Affiliation: