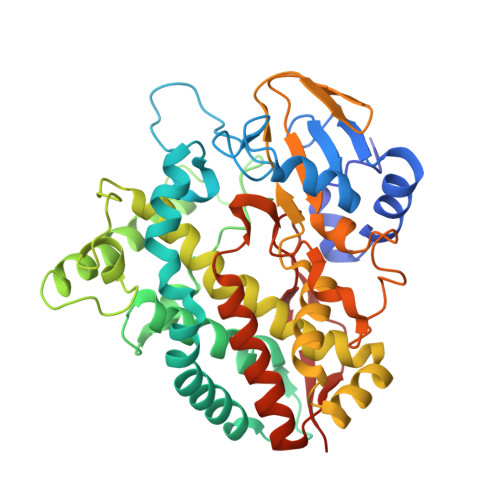
Crystal Structure of a Putative Cytochrome P450 Alkane Hydroxylase (CYP153D17) from Sphingomonas sp. PAMC 26605 and Its Conformational Substrate Binding
Lee, C.W., Yu, S.C., Lee, J.H., Park, S.H., Park, H., Oh, T.J., Lee, J.H.(2016) Int J Mol Sci 17
- PubMed: 27941697
- DOI: https://doi.org/10.3390/ijms17122067
- Primary Citation of Related Structures:
5H1Z - PubMed Abstract:
Enzymatic alkane hydroxylation reactions are useful for producing pharmaceutical and agricultural chemical intermediates from hydrocarbons. Several cytochrome P450 enzymes catalyze the regio- and stereo-specific hydroxylation of alkanes. We evaluated the substrate binding of a putative CYP alkane hydroxylase (CYP153D17) from the bacterium Sphingomonas sp. PAMC 26605. Substrate affinities to C10-C12 n-alkanes and C10-C14 fatty acids with K d values varied from 0.42 to 0.59 μM. A longer alkane (C12) bound more strongly than a shorter alkane (C10), while shorter fatty acids (C10, capric acid; C12, lauric acid) bound more strongly than a longer fatty acid (C14, myristic acid). These data displayed a broad substrate specificity of CYP153D17, hence it was named as a putative CYP alkane hydroxylase. Moreover, the crystal structure of CYP153D17 was determined at 3.1 Å resolution. This is the first study to provide structural information for the CYP153D family. Structural analysis showed that a co-purified alkane-like compound bound near the active-site heme group. The alkane-like substrate is in the hydrophobic pocket containing Thr74, Met90, Ala175, Ile240, Leu241, Val244, Leu292, Met295, and Phe393. Comparison with other CYP structures suggested that conformational changes in the β1-β2, α3-α4, and α6-α7 connecting loop are important for incorporating the long hydrophobic alkane-like substrate. These results improve the understanding of the catalytic mechanism of CYP153D17 and provide valuable information for future protein engineering studies.
- Unit of Polar Genomics, Korea Polar Research Institute, Incheon 406-840, Korea. justay@kopri.re.kr.
Organizational Affiliation: