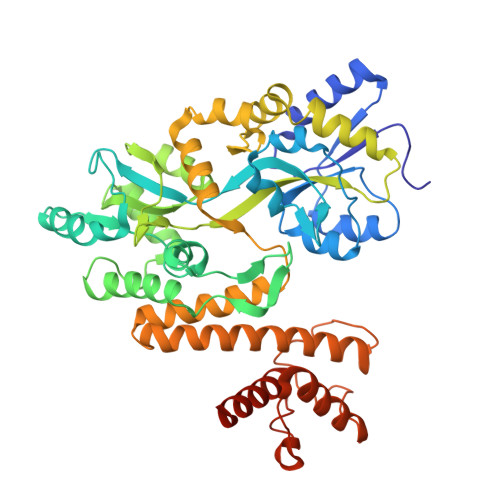
Crystal structure of MBP-PigG fusion protein and the essential function of PigG in the prodigiosin biosynthetic pathway in Serratia marcescens FS14.
Zhang, F., Wei, Q., Tong, H., Xu, D., Wang, W., Ran, T.(2017) Int J Biol Macromol 99: 394-400
- PubMed: 28258005 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2017.02.088
- Primary Citation Related Structures:
5GXT, 5GXV - PubMed Abstract:
Prodigiosin, a tripyrrole red pigment is synthesized by Serratia and some other microbes through a bifurcated biosynthesis pathway; MBC (4-methoxy-2,2'-bipyrrole-5-carbaldehyde) and MAP (2-methyl-3-n-amyl-pyrrole) are synthesized separately and then condensed by PigC to form prodigiosin. PigI, PigG and PigA have been shown to be involved in the first steps of MBC biosynthesis (proline incorporation). The crystal structure of PigG was resolved to elucidate its function and mechanism. PigG, an acyl carrier protein (ACP), features the ACP architecture:, a helical bundle fold containing three major helices and a minor distorted helix together with a conserved "S" motif. An in-frame deletion mutation of the pigG gene abolished the synthesis of prodigiosin in Serratia marcescens FS14. The production of prodigiosin was fully restored by complementation of intact pigG; however the S36A mutant was not able to restore function in the in-frame deletion pigG mutant, indicating that PigG and the conserved serine residue (S36) of PigG are essential for the synthesis of prodigiosin.
- Key Laboratory of Microbiological Engineering of Agricultural Environment, Ministry of Agriculture, Department of Microbiology, College of Life Sciences, Nanjing Agricultural University, Nanjing, China.
Organizational Affiliation: