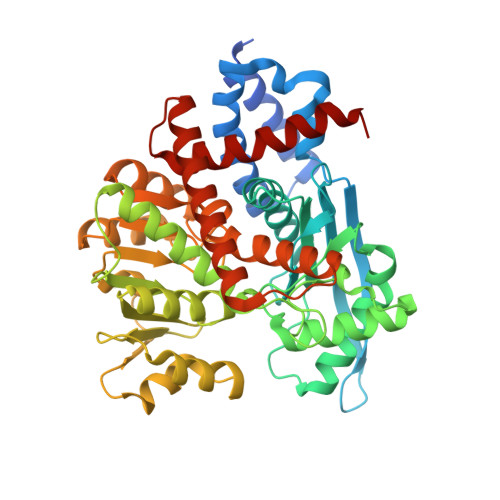
Crystal structure of the 2-iminoglutarate-bound complex of glutamate dehydrogenase from Corynebacterium glutamicum
Tomita, T., Yin, L., Nakamura, S., Kosono, S., Kuzuyama, T., Nishiyama, M.(2017) FEBS Lett 591: 1611-1622
- PubMed: 28486765 Search on PubMed
- DOI: https://doi.org/10.1002/1873-3468.12667
- Primary Citation Related Structures:
5GUD - PubMed Abstract:
The NADP + -dependent glutamate dehydrogenase from Corynebacterium glutamicum (CgGDH) is considered to be one of the key enzymes in the industrial fermentation of glutamate due to its high glutamate-producing activity. We determined the crystal structure of CgGDH complexed with NADP + and 2-iminoglutarate. Among six subunits of hexameric CgGDH-binding NADP + , only four subunits bind 2-iminoglutarate in a closed form, while the other two are in an open form. In the closed form, 2-iminoglutarate is bound to the substrate-binding site with the 2-imino group stacked by the nicotinamide ring of the coenzyme, suggesting a prehydride transfer state in a hypothesized reaction scheme with the imino intermediate. We also conducted MD simulations and provide insights into the extreme preference for the glutamate-producing reaction of CgGDH. The atomic coordinate and structure factors have been deposited in the RCSB PDB database under the accession number 5GUD.
- Biotechnology Research Center, The University of Tokyo, Japan.
Organizational Affiliation: