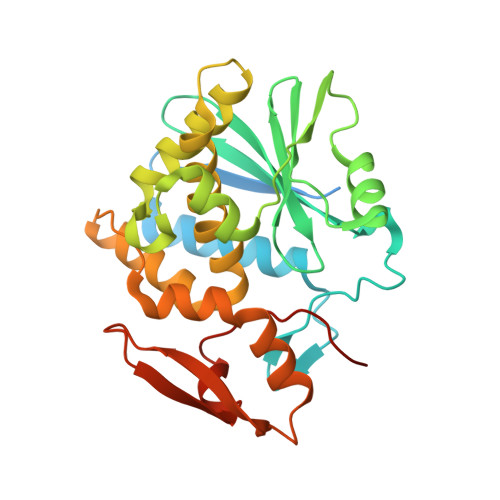
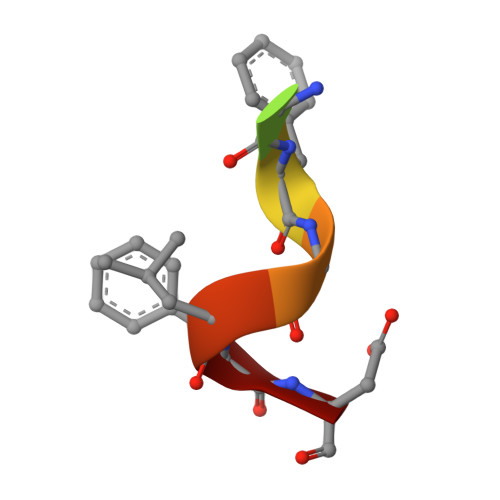
Crystal Structure of Ribosome-Inactivating Protein Ricin A Chain in Complex with the C-Terminal Peptide of the Ribosomal Stalk Protein P2
Shi, W.W., Tang, Y.S., Sze, S.Y., Zhu, Z.N., Wong, K.B., Shaw, P.C.(2016) Toxins (Basel) 8
- PubMed: 27754366 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/toxins8100296
- Primary Citation Related Structures:
5GU4 - PubMed Abstract:
Ricin is a type 2 ribosome-inactivating protein (RIP), containing a catalytic A chain and a lectin-like B chain. It inhibits protein synthesis by depurinating the N-glycosidic bond at α-sarcin/ricin loop (SRL) of the 28S rRNA, which thereby prevents the binding of elongation factors to the GTPase activation center of the ribosome. Here, we present the 1.6 Å crystal structure of Ricin A chain (RTA) complexed to the C-terminal peptide of the ribosomal stalk protein P2, which plays a crucial role in specific recognition of elongation factors and recruitment of eukaryote-specific RIPs to the ribosomes. Our structure reveals that the C-terminal GFGLFD motif of P2 peptide is inserted into a hydrophobic pocket of RTA, while the interaction assays demonstrate the structurally untraced SDDDM motif of P2 peptide contributes to the interaction with RTA. This interaction mode of RTA and P protein is in contrast to that with trichosanthin (TCS), Shiga-toxin (Stx) and the active form of maize RIP (MOD), implying the flexibility of the P2 peptide-RIP interaction, for the latter to gain access to ribosome.
- Centre for Protein Science and Crystallography, School of Life Sciences, The Chinese University of Hong Kong, Shatin, N.T., Hong Kong, China. Shiww@cuhk.edu.hk.
Organizational Affiliation: