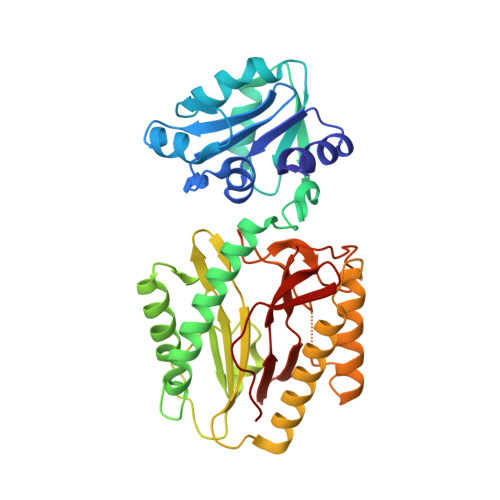
Structures and activities of widely conserved small prokaryotic aminopeptidases-P clarify classification of M24B peptidases.
Are, V.N., Kumar, A., Goyal, V.D., Gotad, S.S., Ghosh, B., Gadre, R., Jamdar, S.N., Makde, R.D.(2018) Proteins
- PubMed: 30536999 Search on PubMed
- DOI: https://doi.org/10.1002/prot.25641
- Primary Citation Related Structures:
5CNX, 5GIQ - PubMed Abstract:
M24B peptidases cleaving Xaa-Pro bond in dipeptides are prolidases whereas those cleaving this bond in longer peptides are aminopeptidases-P. Bacteria have small aminopeptidases-P (36-39 kDa), which are diverged from canonical aminopeptidase-P of Escherichia coli (50 kDa). Structure-function studies of small aminopeptidases-P are lacking. We report crystal structures of small aminopeptidases-P from E. coli and Deinococcus radiodurans, and report substrate-specificities of these proteins and their ortholog from Mycobacterium tuberculosis. These are aminopeptidases-P, structurally close to small prolidases except for absence of dipeptide-selectivity loop. We noticed absence of this loop and conserved arginine in canonical archaeal prolidase (Maher et al., Biochemistry. 43, 2004, 2771-2783) and questioned its classification. Our enzymatic assays show that this enzyme is an aminopeptidase-P. Further, our mutagenesis studies illuminate importance of DXRY sequence motif in bacterial small aminopeptidases-P and suggest common evolutionary origin with human XPNPEP1/XPNPEP2. Our analyses reveal sequence/structural features distinguishing small aminopeptidases-P from other M24B peptidases.
- High Pressure and Synchrotron Radiation Physics Division, Bhabha Atomic Research Centre, Mumbai, Maharashtra, India.
Organizational Affiliation: