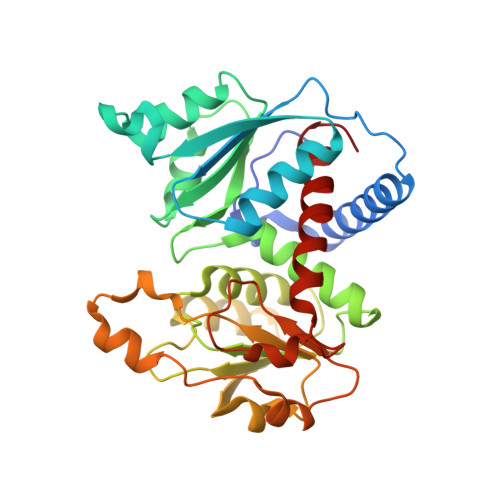
Structure and Functional Characterization of Human Aspartate Transcarbamoylase, the Target of the Anti-Tumoral Drug Pala.
Ruiz-Ramos, A., Velazquez-Campoy, A., Grande-Garcia, A., Moreno-Morcillo, M., Ramon-Maiques, S.(2016) Structure 24: 1081
- PubMed: 27265852 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2016.05.001
- Primary Citation Related Structures:
5G1N, 5G1O, 5G1P - PubMed Abstract:
CAD, the multienzymatic protein that initiates and controls de novo synthesis of pyrimidines in animals, associates through its aspartate transcarbamoylase (ATCase) domain into particles of 1.5 MDa. Despite numerous structures of prokaryotic ATCases, we lack structural information on the ATCase domain of CAD. Here, we report the structure and functional characterization of human ATCase, confirming the overall similarity with bacterial homologs. Unexpectedly, human ATCase exhibits cooperativity effects that reduce the affinity for the anti-tumoral drug PALA. Combining structural, mutagenic, and biochemical analysis, we identified key elements for the necessary regulation and transmission of conformational changes leading to cooperativity between subunits. Mutation of one of these elements, R2024, was recently found to cause the first non-lethal CAD deficit. We reproduced this mutation in human ATCase and measured its effect, demonstrating that this arginine is part of a molecular switch that regulates the equilibrium between low- and high-affinity states for the ligands.
- Structural Bases of Genome Integrity Group, Structural Biology and Biocomputing Programme, Spanish National Cancer Research Centre (CNIO), Melchor Fdez. Almagro, 3, Madrid 28029, Spain.
Organizational Affiliation: